2020-02-04 13:21:25
László Kozák1,2 & Zoltán Szilágyi2 & László Tóth2 & István Pócsi1 & István Molnár
Introduction
Indole-diterpenes (IDTs) are small molecule secondary metabolite natural products biosynthesized by a select group of as- comycetous fungi including Aspergillus, Penicillium, Emericella, Eupenicillium, Claviceps, Epichloë, Escovopsis, Neotyphodium, Periglandula and Tolypocladium species, as well as the zygomycetous fungus Mucor irregularis (Saikia et al. 2008; Schardl et al. 2013; Gao et al. 2016; Dhodary et al. 2018; Supplementary Table S1). IDTs are important myco- toxins that provoke potent neurotoxic and tremorgenic symp- toms in insects and mammals, at least partly due to their inhi- bition of potassium ion channels in the nervous system (Dowd et al. 1988; Uhlig et al. 2009; Imlach et al. 2011). In their native ecological contexts, IDTs defend the overwintering structures of the producing fungi, and serve as effector mole- cules for mutualistic interactions between these fungi and their plant hosts by deterring grazing by large animals and insects (Panaccione et al. 2006; di Menna et al. 2012; Thom et al. 2014). IDT toxicoses of livestock cause significant economic losses in agriculture (Botha et al. 1996; Cawdell-Smith et al. 2010; Philippe 2016). Conversely, IDTs have been considered for the development of pesticides or plant-protecting antifeedants, to be used as components of integrated pest man- agement systems (Panaccione et al. 2014; Saikkonen et al. 2016). Due to their additional, potent and varied bioactivities, IDTs may also serve in the future as drug lead compounds for the development of human and veterinary medications.
IDTs are biosynthesized from geranylgeranyl diphosphate (GGPP) and an indole moiety that originates from a tryptophan precursor (Laws and Mantle 1989; Byrne et al. 2002). After the cloning of the paxilline biosynthetic gene cluster from Penicillium paxilli (Young et al. 2001), the biosynthesis of all the major IDT subgroups were also elucidated from diverse filamentous fungi (Zhang et al. 2004; Young et al. 2006; Motoyama et al. 2012; Nicholson et al. 2015; Kozák et al. 2018). These biosynthetic gene clusters share a common, con- served set of core genes, and are supplemented with additional genes that encode enzymes for various tailoring reactions re- sponsible for the idiosyncratic structural elements of the IDT subgroups and the individual IDT congeners (Zhang et al. 2004; Young et al. 2005; Young et al. 2006; Nicholson et al. 2009). In addition, most IDT biosynthetic enzymes show various levels of substrate and product flexibilities, with even the core set of genes differing in their precise characteristics. Together, these variations create a metabolic grid responsible for the remarkable structural diversity of IDTs produced by fungi.
Our knowledge on the regulation of IDT biosynthesis is still limited. Co-regulation of aflatrem production with sclero- tia development has been demonstrated in A. flavus (Ehrlich and Mack 2014), while a wide spectrum of environmental conditions including temperature, light, various carbon and nitrogen sources were shown to influence penitrem production in Penicillium crustosum (Kalinina et al. 2017). Nevertheless, a deeper understanding of mycotoxin production, including the regulation of the biosynthesis of tremorgenic IDTs will be crucial to combat the agricultural threats posed by these fungi (Uhlig et al. 2009; Moyano et al. 2010; Lee et al. 2017). Moreover, further studies are necessary to estimate the real dimensions of the risks posed by these fungi and their harmful metabolites to consumers of agricultural products (Moldes- Anaya et al. 2012; Eriksen et al. 2013). In particular, the co-occurrence and potential synergistic effects of IDTs with other mycotoxins in foods and feeds warrants further investigations (EFSA Panel on Contaminants in the Food Chain (CONTAM) 2012).
Gaining a deeper insight into the evolution, organization and transcriptional regulation of IDT biosynthetic gene clus- ters may also provide us with valuable tools to control or even eliminate IDT production during industrial fermentations with fungal species such as Claviceps paspali, where the presence of these metabolites represents a safety risk (Kozák et al. 2018). Intriguingly, some paspaline-derived IDTs such as penitrem A may also be regarded as Janus-faced compounds with potential applications in anticancer chemotherapies, ei- ther as monotherapies or in combination with other antiprolif- erative drugs (Sallam et al. 2013a; Sallam et al. 2013b; Goda et al. 2018). Therefore, future metabolic engineering and fer- mentation optimization studies may need to target the im- provement of the yields of these compounds under industrial fermentation conditions (Motoyama et al. 2012; Kalinina et al. 2017). The construction of various fungal synthetic biology platforms for the heterologous production of IDTs with poten- tial biomedical significance is also on the agenda (Tagami et al. 2014; Liu et al. 2015; Tang et al. 2015; Oikawa et al. 2016; Liu et al. 2016).
To the best of our knowledge, this is the first comprehen- sive review that covers the genetic, biochemical, ecological, veterinary, medical and industrial aspects of the production of these important metabolites by fungi.
Detection and structural characterization of IDTs in biological samples
Although the connection between moldy food and some dis- eases such as ergotism was suspected for centuries, it was only in the twentieth century that the occurrence of secondary me- tabolite mycotoxins in these foods was discovered to be the molecular basis for such illnesses (Uraguchi 1969). Recent developments of analytical techniques make it increasingly straightforward to detect, even in an untargeted fashion, a large variety of mycotoxins such as IDTs in complex matrices such as food and feed.
Paxilline and paspaline, the simplest IDT congeners, were first isolated half a century ago and characterized by elemental analysis and various spectroscopic methods such as infrared (IR), ultraviolet-visible (UV-Vis), and mass spectroscopy (MS) (Fehr and Acklin 1966; Cole et al. 1974). Although the first nuclear magnetic resonance (NMR) spectroscopy re- sults about paspaline were published in 1977, the assignments of the signals had to be refined almost two decades later (Munday-Finch et al. 1996). Finally, the complete structures of these mycotoxins were elucidated in 1980 by X-ray crys- tallography (Springer and Clardy 1980).
The discovery of novel IDT subclasses was a relatively slow process in the 1980s, mainly because of the lack of ver- satile analytical techniques. Immunochemical methods like ELISA were considered the most selective and sensitive ana- lytical tools of the era. However, the development of these methods required the very molecule that needed to be ana- lyzed, making these techniques unsuitable for discovery stud- ies. Thin layer chromatography (TLC) served as the most important tool for both the isolation of unknown molecules and the identification of known IDTs (El-Banna et al. 1987; Sanchis et al. 1988; Scuteri et al. 1992). In addition to TLC, high performance liquid chromatography (HPLC) became in- creasingly important in this decade for the analysis of organic compounds, including natural product mycotoxins (Maes et al. 1982; Frisvad 1987; Russell et al. 1989). By 1987 a standardized HPLC method was developed that allowed the detection of all the important groups of mycotoxins and many other fungal secondary metabolites, covering 182 compounds (Frisvad and Thrane 1987). However, the utility of HPLC separation combined with fluorescent detection is rather lim- ited for paspaline-derived IDTs, since only the janthitrems exhibit significant fluorescence (Gallagher et al. 1980b; Lauren and Gallagher 1982). The most traditional mass spec- trometric (MS) technique, electron ionization (EI) was also applied for the analysis of mycotoxins. However, this tech- nique is best suited for volatile compounds (Fellows et al. 1981). Although X-ray crystallography provides detailed in- formation about the structures of organic molecules, it has only been rarely applied to study IDTs (Gallagher et al. 1980a; Nozawa et al. 1987; Kawai and Nozawa 1989) since it requires special pretreatments and relatively larger amounts of the sample.
NMR spectroscopic techniques have gone through a remarkable development in the last few decades, resulting in applications such as 2D NMR spectroscopy (Aue et al. 1976) and magnetic resonance imaging (MRI) (Lauterbur 1973). Research towards the structural elucidation of nov- el IDTs benefited largely from the various 2D NMR tech- niques (De Jesus et al. 1981; Laakso et al. 1992; Wilkins et al. 1992; Belofsky et al. 1995; Munday-Finch et al. 1995). Although examples of HPLC-coupled NMR appli- cations have been described (Sumarah et al. 2005), these methods have not gained widespread acceptance due to the high cost of the instruments and the lower sensitivity of detection as compared to MS.
The most versatile analytical method for the trace analysis of organic substances is mass spectrometry (MS). This is due to the high sensitivity of MS and its ability to provide structural information such as frag- mentation patterns recorded in MS/MS experiments, and elemental composition determined via exact molecular mass measurement with high-resolution MS instruments (Q-TOF MS, Orbitrap, etc.). Since the advent of liquid- phase MS interfaces, most importantly the electrospray ion sources, MS analysis can readily be coupled with HPLC allowing sensitive and selective analysis of com- plex mixtures by measuring directly the molecular mass (typically the mass/charge ratio [m/z] of protonated mol- ecules), including those of new compounds (Naik et al. 1995). The capabilities of the liquid chromatography – mass spectrometry (LC-MS) technique are clearly dem- onstrated by the rapidly increasing number of target bacterial and fungal metabolites that can be analyzed in a single assay (Sulyok et al. 2006; Sulyok et al. 2007; Vishwanath et al. 2009; Malachova et al. 2014). In addition to multicomponent analysis, the most ad- vanced high-resolution MS approaches allow researchers to perform untargeted LC-MS surveys, resulting in a complex dataset containing information about the mo- lecular masses (normal MS spectra) and about the struc- tures (fragmentation and isotopic patterns) represented in the analyte. The collected datasets can be retrospectively examined even years after the actual measurement, with- out the need to repeat the experiment (Andersen et al. 2016; Renaud and Sumarah 2016).
Biosynthesis of paspaline-derived IDTs in fungi
Biosynthesis of paspaline
Paspaline is the founding member of the paspaline- derived IDTs that feature an angular hexacyclic ring sys- tem consisting of a tetracyclic diterpene fused to an indole moiety (Fig. 1). This scaffold is further elaborated by prenylation, oxidation, reduction, cyclization and chlori- nation in the various members of the group (Saikia et al. 2007; Tagami et al. 2013; Liu et al. 2016). Paspaline- derived IDTs have been classified in various ways, using historic and structural criteria. Parker et al. suggested nine subclasses: the penitrems, janthitrems, lolitrems, aflatrems, terpendoles, shearinines, sulpinines, paxilline and the paspaline/paspalinine/paspalitrems group (Parker and Scott 2005). Another classification by Sings et al. divides IDTs into six structural groups: paspalanes, aflatremanes, penitremanes, janthitremanes, lolitremanes and nodulisporanes (Sings and Singh 2003). In Fig. 2, we classified paspaline-derived IDTs into three main sub- groups (paxilline-type, paspalinine-type and terpendole- type compounds) according to their diverging biosynthet- ic pathways. Nodulisporanes (Sings and Singh 2003) are not derived from paspaline, and lack the F ring of the paspaline skeleton (Fig. 1), so they are not discussed in this review.
The cyclic diterpene moiety of paspaline is derived from geranylgeranyl diphosphate (GGPP), biosynthesized from farnesyl diphosphate by the GGPP synthase PaxG in
P. paxilli and its orthologues in other IDT producers (Fig. 1) (Tagami et al. 2013). Deletion of paxG results in the complete elimination of the production of the whole spectrum of IDTs in P. paxilli (Young et al. 2001; Saikia and Scott 2009). Orthologues of paxG were detected in all characterized IDT gene clusters except that of the terpendole K producer Tolypocladium album (Motoyama et al. 2012).
The indole group of IDTs most likely derives from indole- 3-glycerol phosphate (Fig. 1). Prenylation of C3 of the indole with the concomitant elimination of glyceraldehyde 3- phosphate yields 3-geranylgeranyl indole (GGI), a common intermediate for all IDTs. This reaction is catalyzed by the prenyl transferase PaxC in P. paxilli and its orthologues in other fungi. Although the preferred substrate for prenylation is indole-3-glycerol phosphate, tryptophan was also accept- ed by recombinant PaxC during in vitro studies with the pu- rified enzyme (Tagami et al. 2013). Epoxidation of GGI by PaxM, a FAD-dependent monooxygenase (or its orthologues) yields 10(11)-epoxygeranylgeranyl indole. Cyclization of this intermediate by PaxB or its orthologues affords emindole SB. Finally, another epoxidation cyclization sequence, catalyzed again by the PaxM - PaxB pair and their orthologues in other fungi, yields paspaline through the formation of the F ring of the IDT skeleton (Tagami et al. 2013; Van de Bittner et al. 2018).
Generation of chemical diversity in the paspaline-derived IDT families
IDT biosynthesis diverges after paspaline, with further modi- fications of this common intermediate by various oxidations, reductions and prenylations leading to the amazing chemical diversity of the various IDT families (Fig. 2). The first diver- gent reaction is the oxidation of paspaline. On the first branch, TerQ-catalyzed hydroxylation of paspaline at C11 yields terpendole E in T. album (Motoyama et al. 2012). This inter- mediate is further oxidized by TerP to eliminate the pendant methyl group on C12, yielding 13-desoxyterpendole I that features a C11(12) epoxide and a C10 alcohol (Motoyama et al. 2012). The C11(12) epoxide is also present in lolitrem B produced by Neotyphodium lolii, suggesting that the TerQ orthologue LtmQ also possesses C11 hydroxylation activity (Gallagher et al. 1981; Philippe 2016). Correspondingly, the terpendole E-like compound lolicine was isolated from the lolitrem producer N. lolii (Munday-Finch et al. 1998). Next, TerQ catalyzes another oxidation, this time the hydroxylation of C13 to form terpendole I. This intermediate is O-prenylated at C27 by TerF or its orthologues such as LtmF of N. lolii or Epichloë festucae. Next, oxidative acetal ring formation by the cytochrome P450 TerK or LtmK gives rise to terpendole C. Additional diprenylation at C20 and C21 by LtmE and oxida- tive ring closure by the cytochrome P450 LtmJ of N. lolii provide the lolitremane IDTs in N. lolii/E. festucae (Young et al. 2006; Saikia et al. 2012).
On the other branch diverging from paspaline (Fig. 2), the cytochrome P450 monooxygenase PaxP and its orthologues catalyze the oxidative elimination of the pendant methyl group connected to C12 and the formation of the C10 ketone to yield 13-desoxypaxilline via the intermediate β-PC-M6 (McMillan et al. 2003). From 13-desoxypaxilline, the pathway again bi- furcates. PaxQ-catalyzed hydroxylation of C13 in 13- desoxypaxilline yields paxilline en route to the pentirem/ jantithrem/sulpinine families of IDTs (McMillan et al. 2003; Nicholson et al. 2015). On the other branch, AtmQ of A. flavus and its orthologues such as IdtQ of C. paspali also hydroxyl- ate C13, but they oxidize C7 as well to form a cyclic acetal with the C27 alcohol to afford paspalinine. The order of the C13 vs. C7 oxidations is still unclear and may even be dissim- ilar in different fungi (Nicholson et al. 2009; Kozák et al. 2018). Thus, during the elucidation of the IDT product spec- trum of C. paspali, paxilline and paspalinine were both de- tected in sclerotia extracts together with paspalicine, the 13- desoxy analogue of paspalinine. This indicates that IdtQ can accept 13-desoxypaxilline for either C7 or C13 oxidation in this fungus (Uhlig et al. 2014). On the other hand, paspalicine could not be isolated during the heterologous production of aflatrems using AtmQ of A. flavus, suggesting that C13 hy- droxylation of 13-desoxypaxilline may precede the formation of the cyclic acetal with that enzyme (Nicholson et al. 2009).
In any case, formation of paspalinine opens the way towards the biosynthesis of the aflatrems, paspalitrems and shearinines. Thus, the substrate and product specificities of the TerQ/PaxQ/AtmQ-like and the TerP/PaxP/AtmP-like cy- tochrome P450 enzymes in different fungi define the F ring architecture of the IDT skeleton, resulting in the terpendole/ lolitrem, the penitrem/janthitrem/sulpinine, and the paspalitrem/aflatrem/shearinine subgroups of paspaline- derived IDTs.
Aflatrem, β-aflatrem, and the paspalitrems are the monoprenylated derivatives of paspalinine (Fig. 2). In the case of aflatrems, the dimethylallyl transferase AtmD catalyzes the γ-selective prenylation of paspalinine at the C20 or the C21 positions to afford aflatrem A and β-aflatrem, respectively. Interestingly, AtmD also accepts paxilline and paspaline as substrates (Liu et al. 2013). While paxilline is prenylated at C20 or C21 just as in paspalinine, paspaline is modified at the C21 or C22 positions. For paspalitrems, the IdtF dimethylallyl transferase of C. paspali conducts an α-selective prenylation of paspalinine at the C21 or the C20 positions to yield paspalitrem A or paspalitrem C, respectively. Paspalitrem A is then further oxidized at C32 giving rise to paspalitrem B. For shearinines, the JanD dimethylallyl transferase from P. janthinellum conducts two consecutive α-selective prenylations at C21 and C22 to yield shearinine K (Nicholson et al. 2015; Liu et al. 2016). This intermediate undergoes a two-step oxidative cyclization sequence cata- lyzed by the JanO FAD-dependent oxidase that yields a bicy- clic ring system fused to the paspalinine core, affording shearinine A. Finally, an additional hydroxylation conducted by the cytochrome P450 JanJ gives rise to shearinine D.
At the paxilline branch of the biosynthetic pathway (Fig. 2), the biosynthesis of penitrems, the most elaborate paspaline-derived IDTs, requires 17 enzymes in P. simplicissimum. Paxilline is first reduced by PtmH at the C10 ketone to yield the corresponding alcohol, followed by α- selective prenylation at C20 by the dimethylallyl transferase PtmD. The prenylated analogue undergoes an acetylation- elimination sequence catalyzed by PtmV and PtmI to yield 20-prenylpenijanthine with a terminal olefin. The order of the PtmH-PtmD-PtmVI reactions seems to be somewhat flex- ible (Oikawa et al. 2016). Biosynthesis of janthitrems may follow a similar sequence, with the exception that a diprenylase-oxidative cyclase pair orthologous to JanD-JanO in shearinine biosynthesis is involved in the formation of the bicyclic system fused to the paxilline ring A. Indeed, the P. paxilli gene cluster contains the JanD and JanO orthologues PaxD and PaxO, respectively. Paxilline is an excellent sub- strate for recombinant PaxD that produces 21,22- diprenylpaxilline. While PaxD and PaxO are silent (or at least very weakly expressed) under normal fermentation conditions thus leaving the fungus to produce paxilline as its major IDT product, the P. paxilli gene cluster may nevertheless encode the biosynthesis of a janthitrem congener (Liu et al. 2016). Similarly, sulpinine biosynthesis may follow a sequence anal- ogous to that of PtmH-PtmD-PtmVI for penitrems, with the relevant dimethylallyl transferase catalyzing a γ-selective prenylation at C22 on ring A.
For penitrems, γ-hydroxylation of the prenyl side chain by PtmO prefaces a head-to-head coupling with dimethylallyl diphosphate during a prenylation-initiated cationic cyclization event catalyzed by the PtmE dimethylallyl transferase. Oxidative ring expansion by the PtmK cytochrome P450 yields the bicyclo[4,2,0]octane system, followed by the for- mation of the 8-membered oxocane cyclic ether bridging to C17, catalyzed by another cytochrome P450 (PtmU). Epoxide formation at C11(12) by PtmL, chlorination at C22, and hy- droxylation of the bicyclooctane by PtmJ finally affords penitrem A (Liu et al. 2015).
It is important to note that many of the IDT biosynthetic enzymes described here display substrate and product flexi- bility, leading to a significant metabolic crosstalk among the pathways leading to the three main biogenetic subgroups of IDTs. Consequently, many fungi use a single set of enzymes that form a complex biosynthetic grid with considerable plas- ticity, thus producing multiple paspaline-derived metabolites that belong to more than one IDT chemical families. Similarly, the structural classification of paspaline-derived IDTs is also somewhat arbitrary and depends on the structural elements or scaffold tailoring events that are considered the most impor- tant by the authors. For example, janthithremanes are often understood to include the janthitrems (derived from paxilline) and the shearinines (originating from paspalinine).
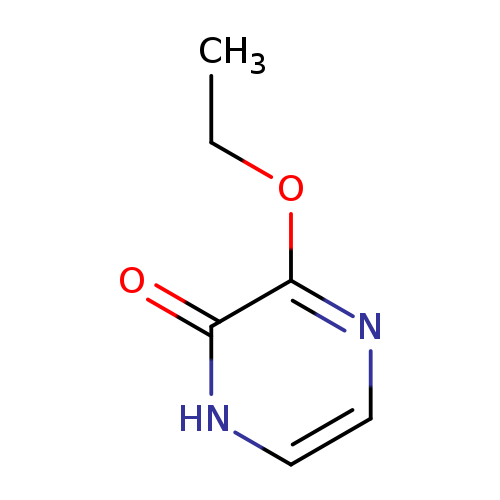
3-Ethoxy-1,2-dihydropyrazin-2-oneCatalog No.:AA01A49C CAS No.:1093656-66-6 MDL No.:MFCD19227814 MF:C6H8N2O2 MW:140.1399 |
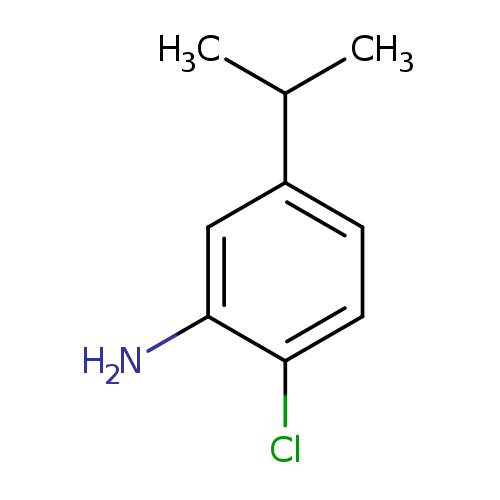
2-chloro-5-(propan-2-yl)anilineCatalog No.:AA01FL1L CAS No.:1093657-08-9 MDL No.:MFCD19598714 MF:C9H12ClN MW:169.6513 |
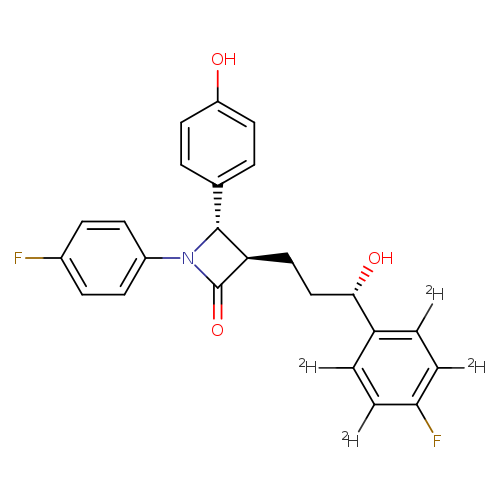
Ezetimibe-d4Catalog No.:AA008X0L CAS No.:1093659-89-2 MDL No.: MF:C24H17D4F2NO3 MW:413.4499 |
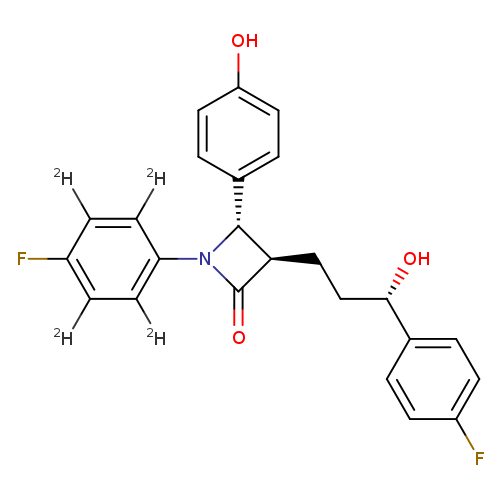
EZETIMIBE-D4Catalog No.:AA008TJ8 CAS No.:1093659-90-5 MDL No.:MFCD08063570 MF:C24H17D4F2NO3 MW:413.4499 |
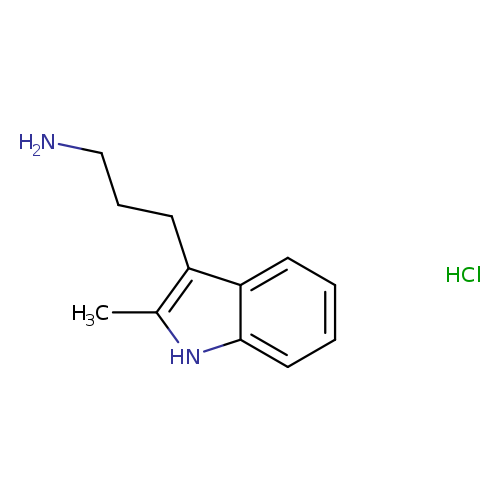
[3-(2-methyl-1H-indol-3-yl)propyl]amine hydrochlorideCatalog No.:AA00J0SM CAS No.:109366-84-9 MDL No.:MFCD13250089 MF:C12H17ClN2 MW:224.7298 |
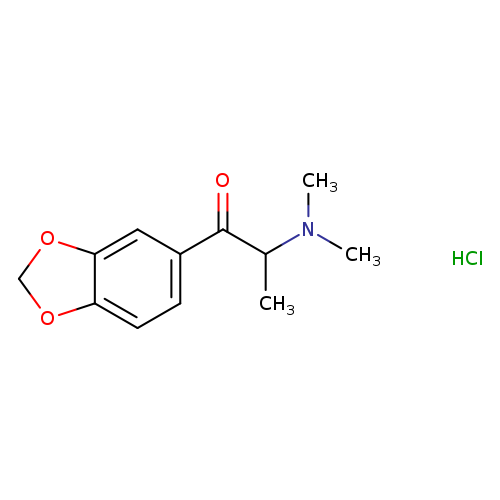
bk-MDDMA (hydrochloride)Catalog No.:AA008SOH CAS No.:109367-07-9 MDL No.:MFCD30718088 MF:C12H16ClNO3 MW:257.7133 |
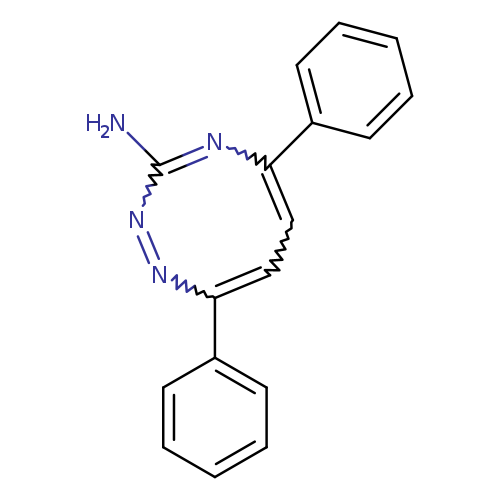
3-AMINO-5,8-DIPHENYL-1,2,4-TRIAZOCINECatalog No.:AA01FOGY CAS No.:109367-42-2 MDL No.:MFCD00014121 MF:C17H14N4 MW:274.3199 |
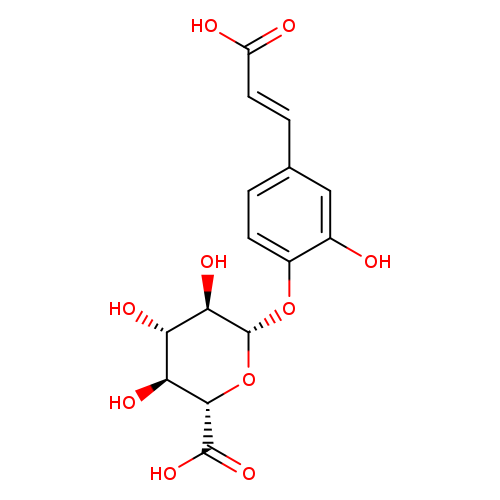
Caffeic Acid 4-β-D-GlucuronideCatalog No.:AA008WEX CAS No.:1093679-71-0 MDL No.:MFCD30186676 MF:C15H16O10 MW:356.2815 |
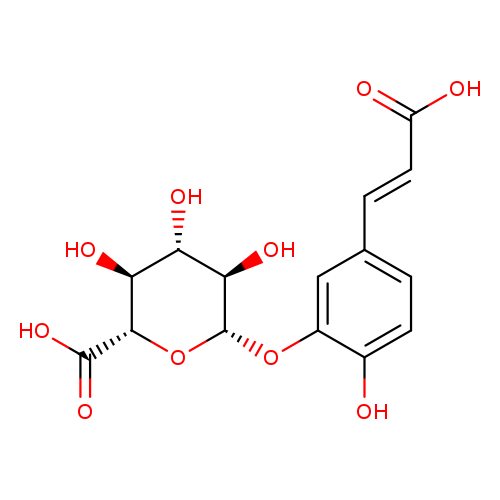
5-(2-Carboxyethenyl)-2-hydroxyphenyl-beta-d-glucopyranosiduronic acidCatalog No.:AA008WBG CAS No.:1093679-73-2 MDL No.:MFCD28122829 MF:C15H16O10 MW:356.2815 |
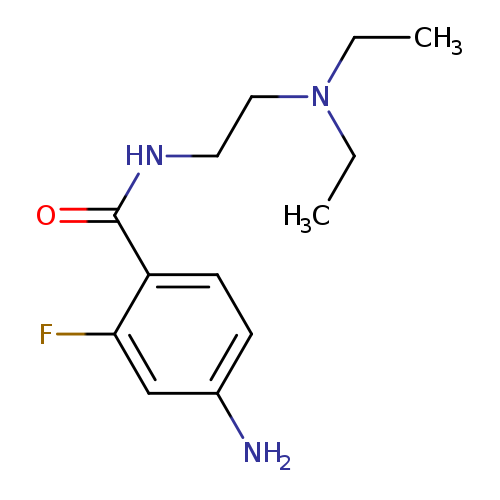
4-amino-N-[2-(diethylamino)ethyl]-2-fluorobenzamideCatalog No.:AA01A0V7 CAS No.:1093728-01-8 MDL No.:MFCD16508390 MF:C13H20FN3O MW:253.3158 |
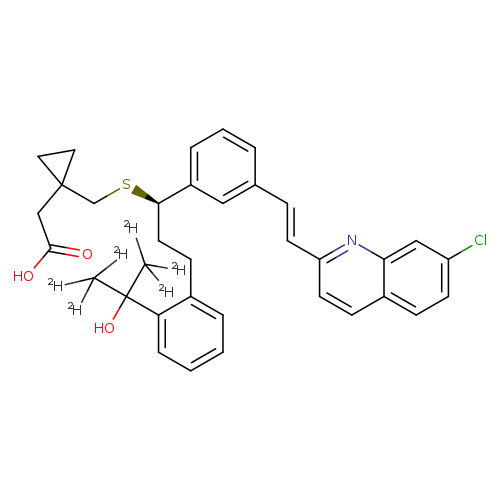
Montelukast-d6Catalog No.:AA008WM4 CAS No.:1093746-29-2 MDL No.:MFCD19705020 MF:C35H30ClD6NO3S MW:592.2202 |
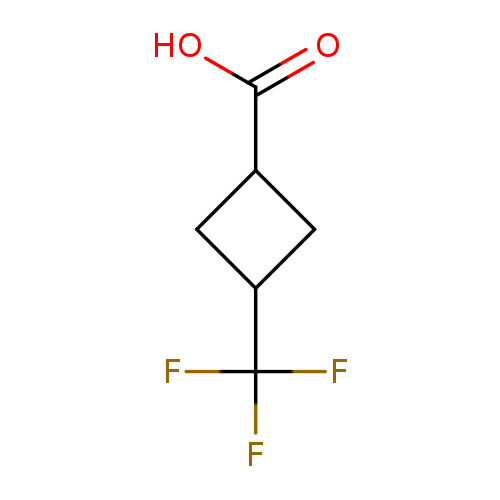
3-(Trifluoromethyl)cyclobutane-1-carboxylic acidCatalog No.:AA00HBD3 CAS No.:1093750-93-6 MDL No.:MFCD15526994 MF:C6H7F3O2 MW:168.1138 |
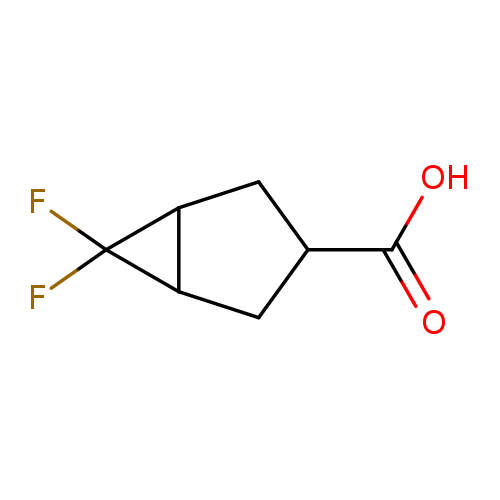
6,6-Difluorobicyclo[3.1.0]hexane-3-carboxylic acidCatalog No.:AA00HBD5 CAS No.:1093751-02-0 MDL No.:MFCD27923654 MF:C7H8F2O2 MW:162.1340 |
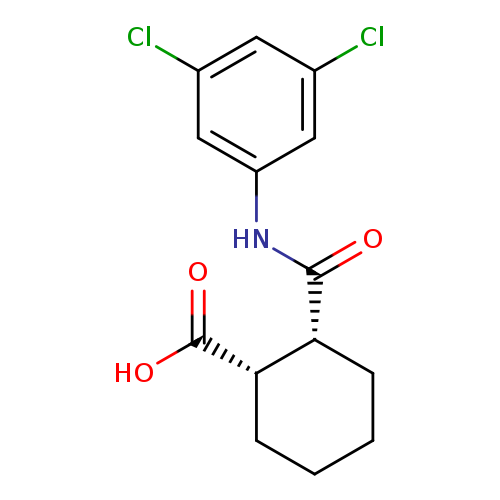
VU0155041Catalog No.:AA008WVA CAS No.:1093757-42-6 MDL No.:MFCD03544581 MF:C14H15Cl2NO3 MW:316.1798 |
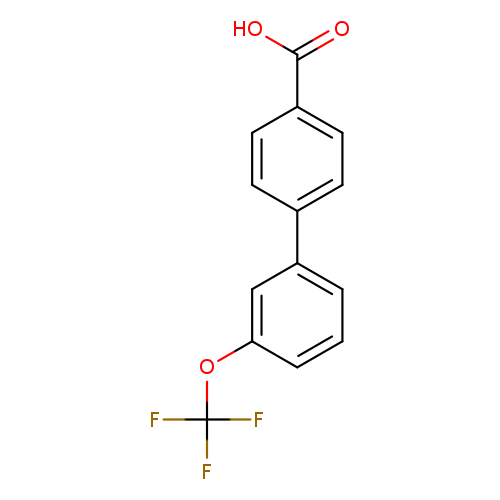
4-(3-Trifluoromethoxyphenyl)benzoic acidCatalog No.:AA007T40 CAS No.:1093758-81-6 MDL No.:MFCD11519023 MF:C14H9F3O3 MW:282.2147 |
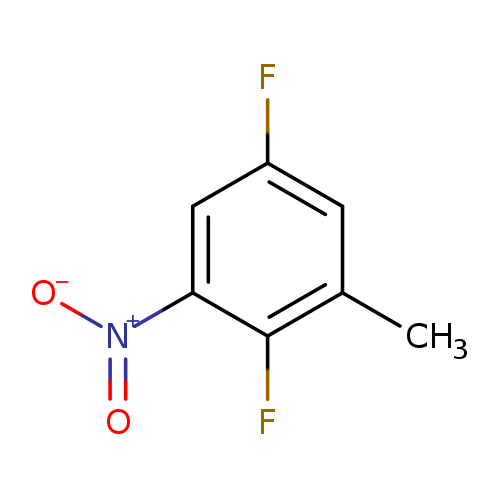
2,5-Difluoro-3-nitrotolueneCatalog No.:AA008UCO CAS No.:1093758-82-7 MDL No.:MFCD11519028 MF:C7H5F2NO2 MW:173.1169 |
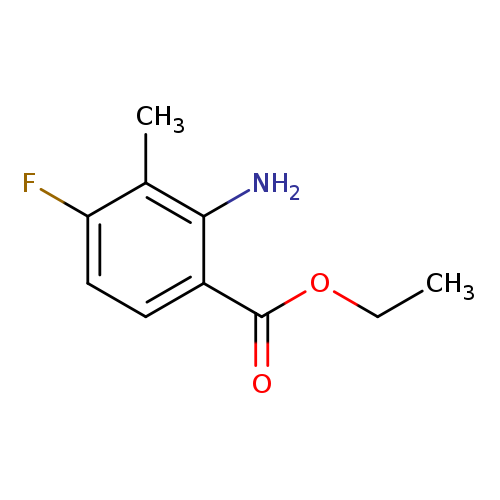
Ethyl 2-amino-4-fluoro-3-methylbenzoateCatalog No.:AA008UED CAS No.:1093758-83-8 MDL No.:MFCD11519034 MF:C10H12FNO2 MW:197.2062 |
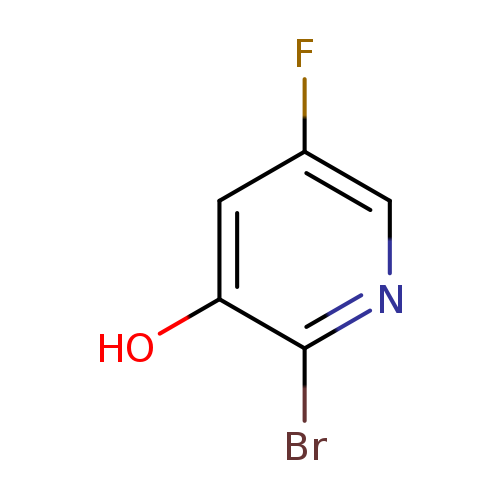
2-Bromo-5-fluoropyridin-3-olCatalog No.:AA008UD6 CAS No.:1093758-87-2 MDL No.:MFCD11519042 MF:C5H3BrFNO MW:191.9858 |
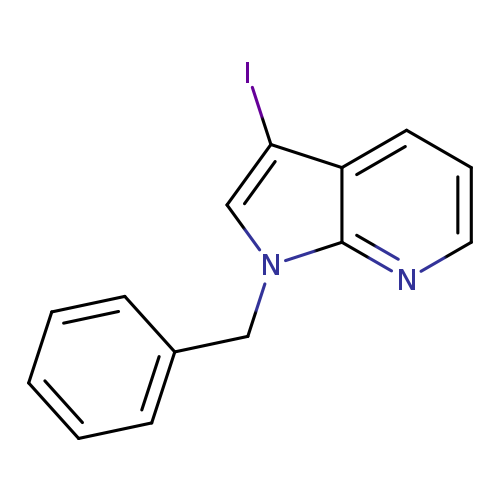
1-Benzyl-3-iodo-7-azaindoleCatalog No.:AA00HBD7 CAS No.:1093759-51-3 MDL No.:MFCD09834873 MF:C14H11IN2 MW:334.1550 |
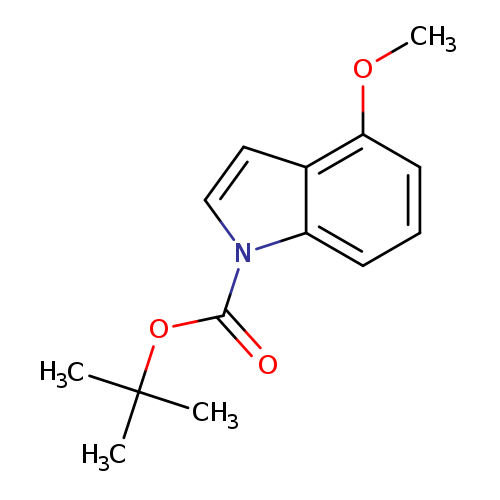
tert-Butyl 4-methoxy-1h-indole-1-carboxylateCatalog No.:AA007AUG CAS No.:1093759-59-1 MDL No.:MFCD09834877 MF:C14H17NO3 MW:247.2897 |
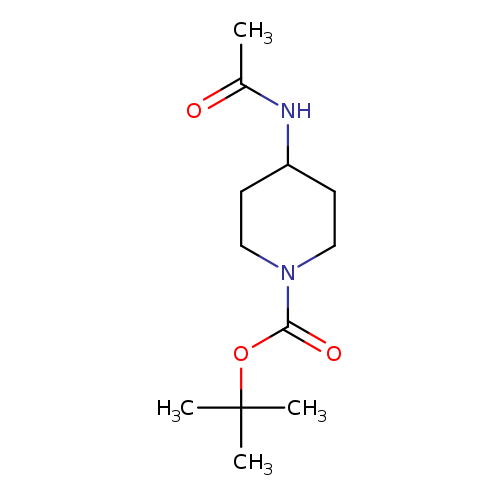
1-BOC-4-(Acetamido)piperidineCatalog No.:AA00HBD8 CAS No.:1093759-67-1 MDL No.:MFCD20482687 MF:C12H22N2O3 MW:242.3147 |

3-Difluoromethylpiperidine HClCatalog No.:AA008TSJ CAS No.:1093759-69-3 MDL No.:MFCD18325187 MF:C6H12ClF2N MW:171.6160 |
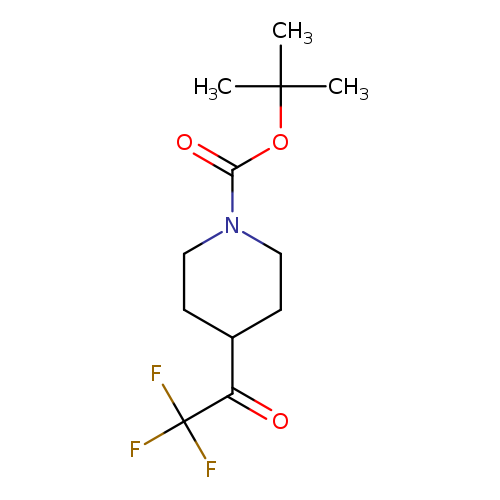
tert-Butyl 4-(2,2,2-trifluoroacetyl)piperidine-1-carboxylateCatalog No.:AA003A0Q CAS No.:1093759-80-8 MDL No.:MFCD11501650 MF:C12H18F3NO3 MW:281.2714 |
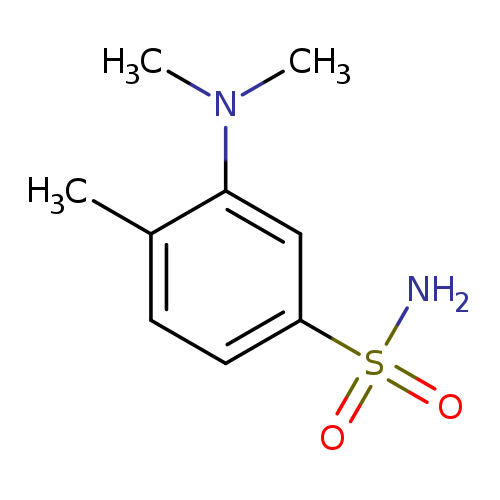
3-(dimethylamino)-4-methylbenzene-1-sulfonamideCatalog No.:AA01A05R CAS No.:1093759-99-9 MDL No.:MFCD22378665 MF:C9H14N2O2S MW:214.2847 |
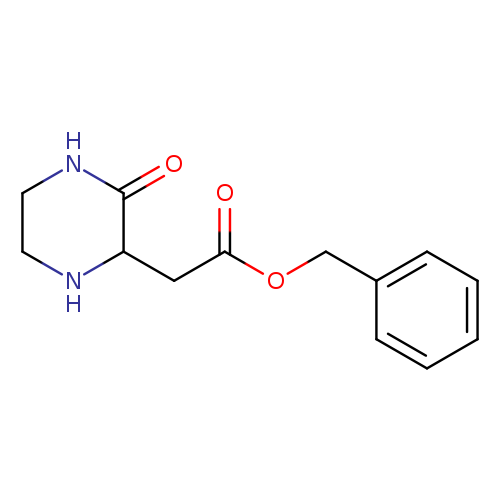
Benzyl 2-(3-oxopiperazin-2-yl)acetateCatalog No.:AA0090T2 CAS No.:1093760-28-1 MDL No.:MFCD05666693 MF:C13H16N2O3 MW:248.2777 |
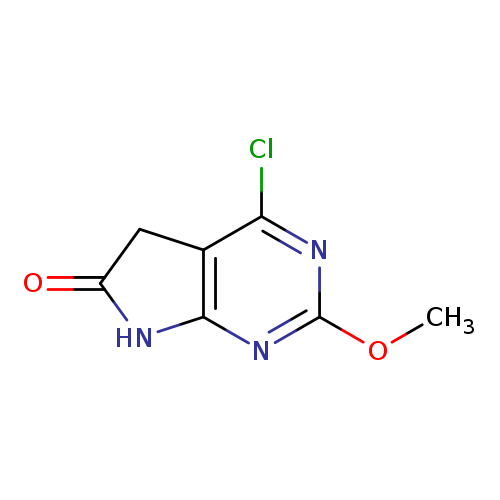
4-chloro-2-methoxy-5H,6H,7H-pyrrolo[2,3-d]pyrimidin-6-oneCatalog No.:AA01DX7R CAS No.:1093799-53-1 MDL No.:MFCD21362713 MF:C7H6ClN3O2 MW:199.5944 |
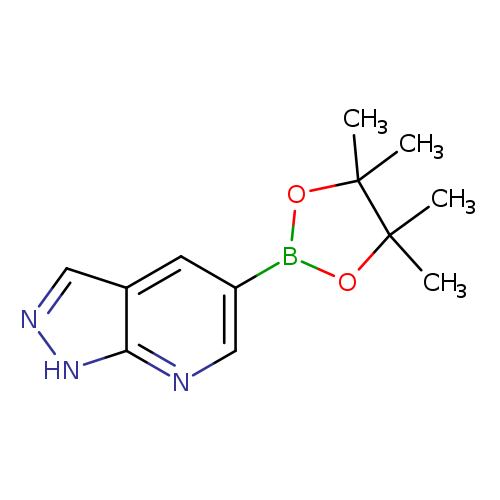
1H-Pyrazolo[3,4-b]pyridine-5-boronic acid pinacol esterCatalog No.:AA0082GK CAS No.:1093819-50-1 MDL No.:MFCD09832892 MF:C12H16BN3O2 MW:245.0853 |
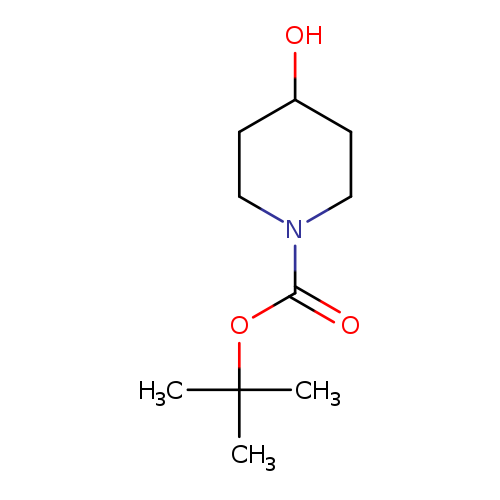
tert-Butyl 4-hydroxypiperidine-1-carboxylateCatalog No.:AA0037VX CAS No.:109384-19-2 MDL No.:MFCD01075174 MF:C10H19NO3 MW:201.2628 |
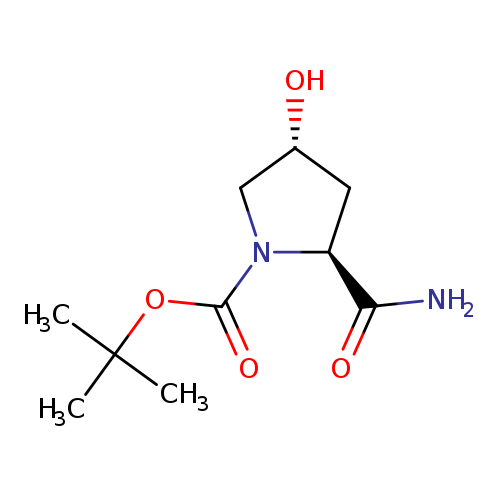
(2S,4R)-1-Boc-2-carbamoyl-4-hydroxypyrrolidineCatalog No.:AA003BMK CAS No.:109384-24-9 MDL No.:MFCD16883081 MF:C10H18N2O4 MW:230.2609 |
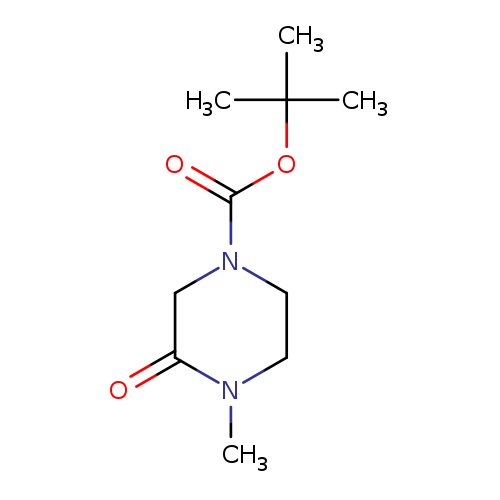
tert-Butyl 4-methyl-3-oxopiperazine-1-carboxylateCatalog No.:AA008UPJ CAS No.:109384-26-1 MDL No.:MFCD10574956 MF:C10H18N2O3 MW:214.2615 |
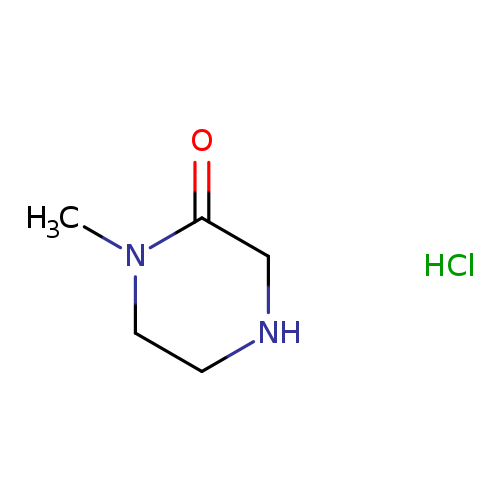
1-Methylpiperazin-2-one hydrochlorideCatalog No.:AA0082DP CAS No.:109384-27-2 MDL No.:MFCD06662557 MF:C5H11ClN2O MW:150.6066 |
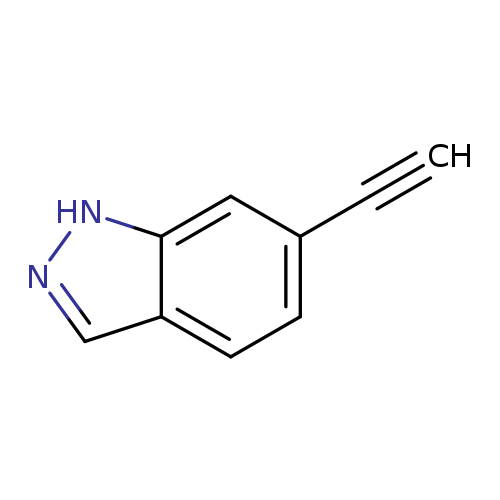
6-Ethynyl-1H-indazoleCatalog No.:AA00HBDE CAS No.:1093847-80-3 MDL No.:MFCD20527779 MF:C9H6N2 MW:142.1573 |
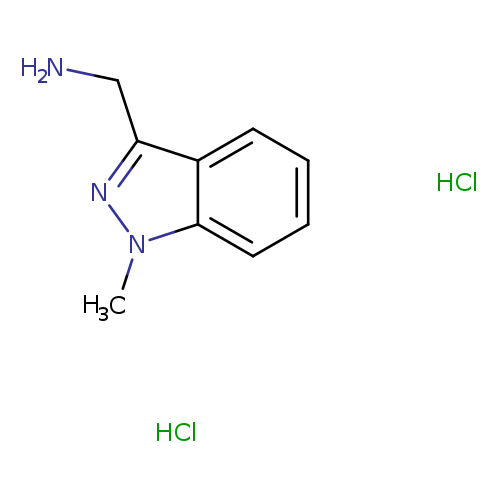
(1-Methyl-1H-indazol-3-yl)methanamine dihydrochlorideCatalog No.:AA0082DN CAS No.:1093860-45-7 MDL No.:MFCD11841065 MF:C9H13Cl2N3 MW:234.1256 |
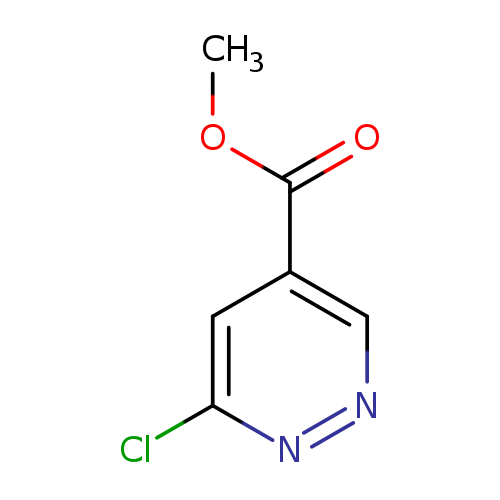
Methyl 6-chloropyridazine-4-carboxylateCatalog No.:AA00355L CAS No.:1093860-48-0 MDL No.:MFCD16660968 MF:C6H5ClN2O2 MW:172.5691 |
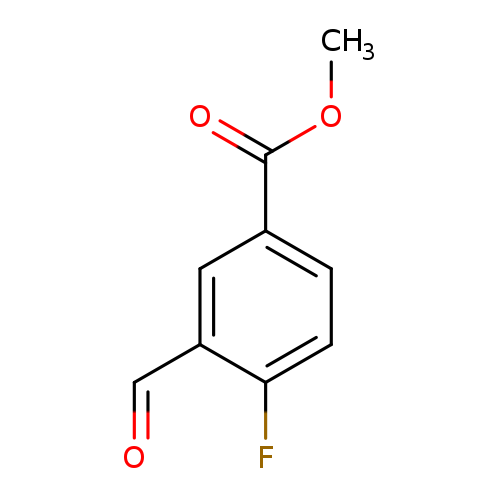
Methyl 4-fluoro-3-formylbenzoateCatalog No.:AA007AUA CAS No.:1093865-65-6 MDL No.:MFCD16036861 MF:C9H7FO3 MW:182.1485 |
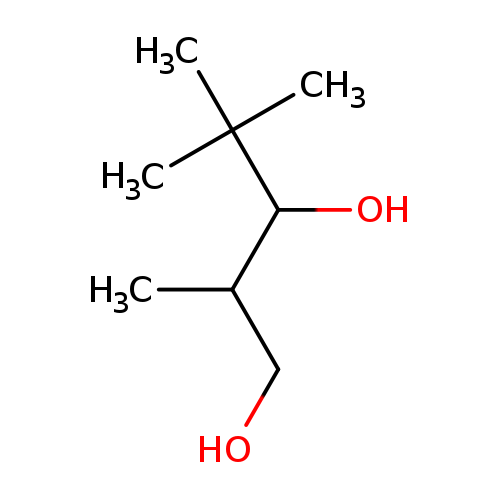
2,4,4-Trimethylpentane-1,3-diolCatalog No.:AA01BRYW CAS No.:109387-36-2 MDL No.:MFCD06252429 MF:C8H18O2 MW:146.2273 |
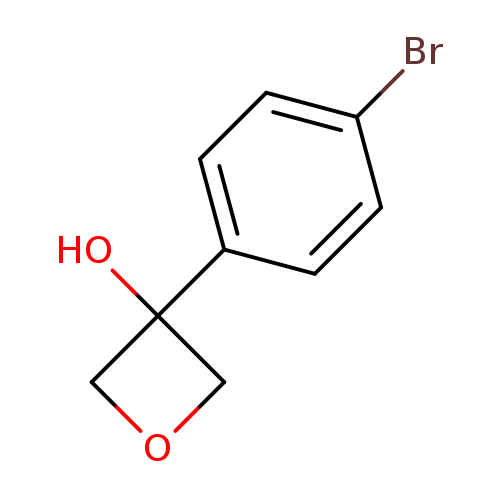
3-(4-Bromophenyl)oxetan-3-olCatalog No.:AA008XMT CAS No.:1093878-32-0 MDL No.:MFCD16657878 MF:C9H9BrO2 MW:229.0706 |
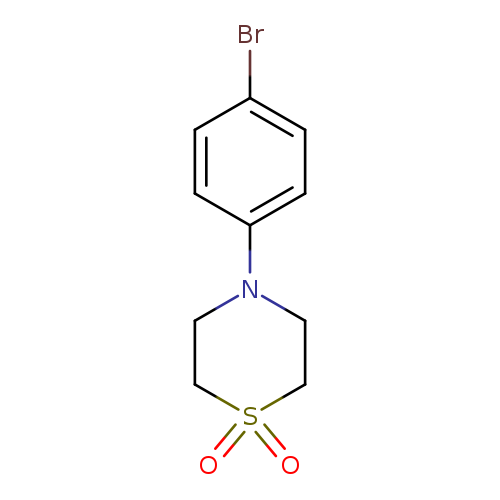
4-(4-bromophenyl)thiomorpholine-1,1-dioneCatalog No.:AA003JZZ CAS No.:1093878-42-2 MDL No.:MFCD08276281 MF:C10H12BrNO2S MW:290.1768 |
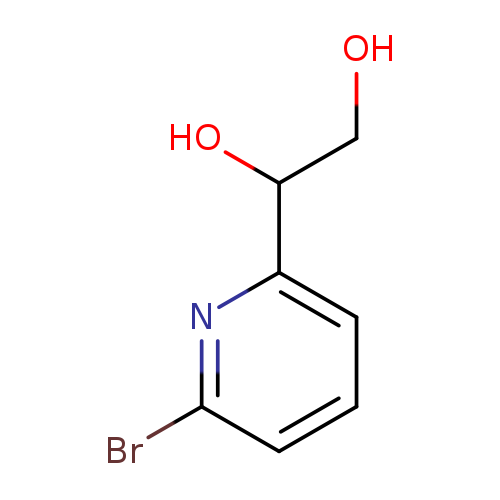
1-(6-bromopyridin-2-yl)ethane-1,2-diolCatalog No.:AA01EL8E CAS No.:1093879-16-3 MDL No.:MFCD28403476 MF:C7H8BrNO2 MW:218.0479 |
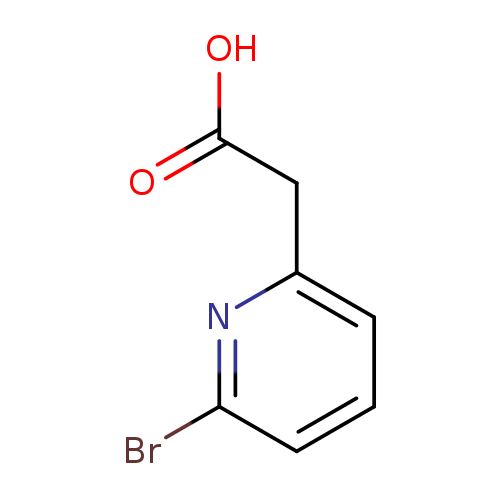
2-(6-Bromopyridin-2-yl)acetic acidCatalog No.:AA008VDO CAS No.:1093879-46-9 MDL No.:MFCD10000075 MF:C7H6BrNO2 MW:216.0320 |
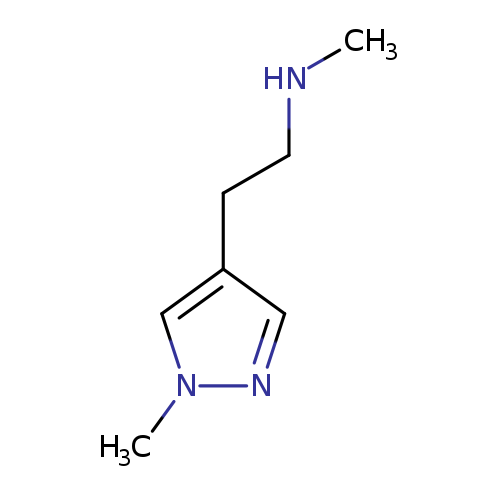
N-Methyl-2-(1-methyl-1h-pyrazol-4-yl)ethanamineCatalog No.:AA008V7K CAS No.:1093879-63-0 MDL No.:MFCD16556159 MF:C7H13N3 MW:139.1982 |
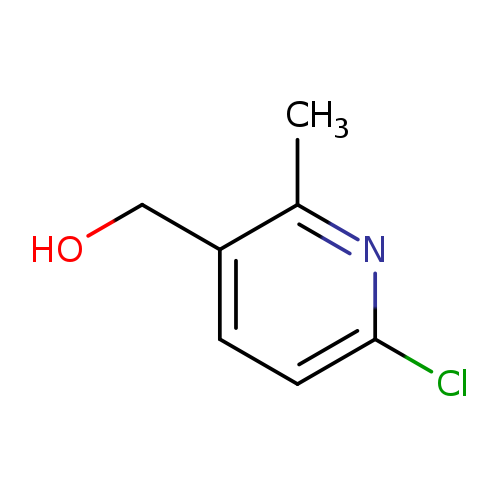
(6-Chloro-2-methylpyridin-3-yl)methanolCatalog No.:AA0096WB CAS No.:1093879-95-8 MDL No.:MFCD13689158 MF:C7H8ClNO MW:157.5975 |
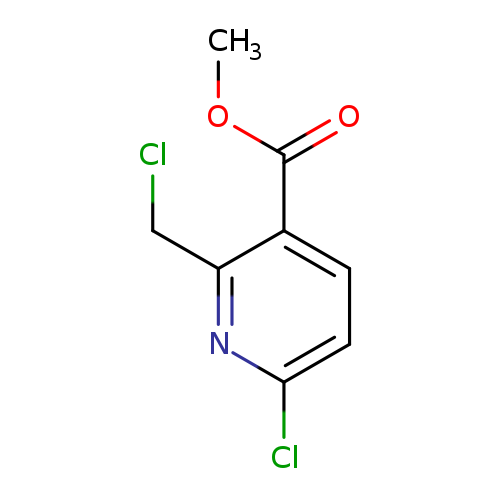
Methyl 6-chloro-2-(chloromethyl)nicotinateCatalog No.:AA00HBDM CAS No.:1093879-99-2 MDL No.:MFCD16658090 MF:C8H7Cl2NO2 MW:220.0527 |
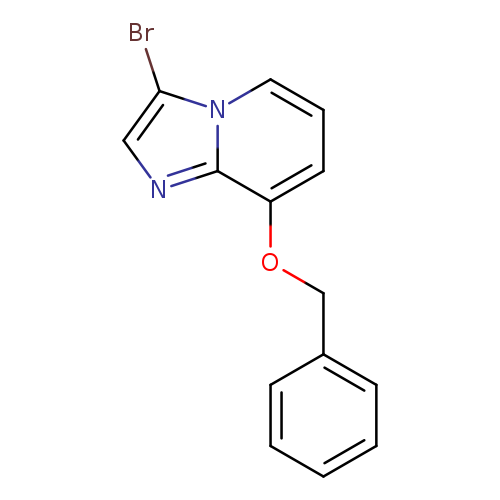
8-(Benzyloxy)-3-bromoimidazo[1,2-a]pyridineCatalog No.:AA0095XY CAS No.:109388-59-2 MDL No.:MFCD21605751 MF:C14H11BrN2O MW:303.1539 |
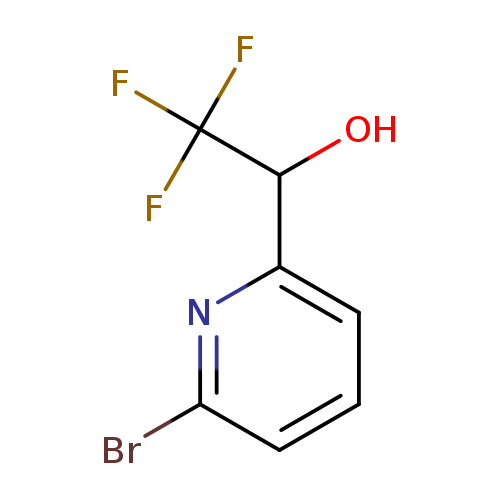
1-(6-bromopyridin-2-yl)-2,2,2-trifluoroethan-1-olCatalog No.:AA01AN3Y CAS No.:1093880-21-7 MDL No.:MFCD16657881 MF:C7H5BrF3NO MW:256.0199 |
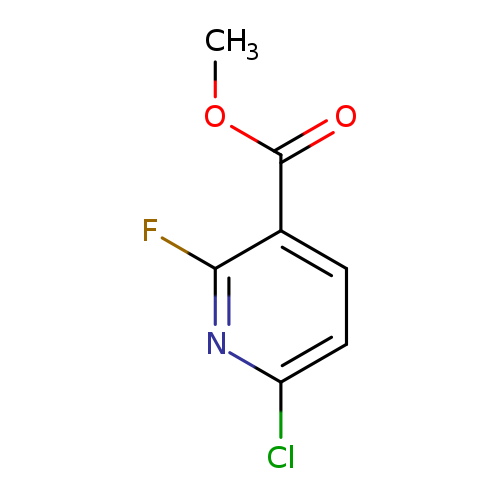
Methyl 6-chloro-2-fluoronicotinateCatalog No.:AA00999C CAS No.:1093880-34-2 MDL No.:MFCD16658093 MF:C7H5ClFNO2 MW:189.5715 |
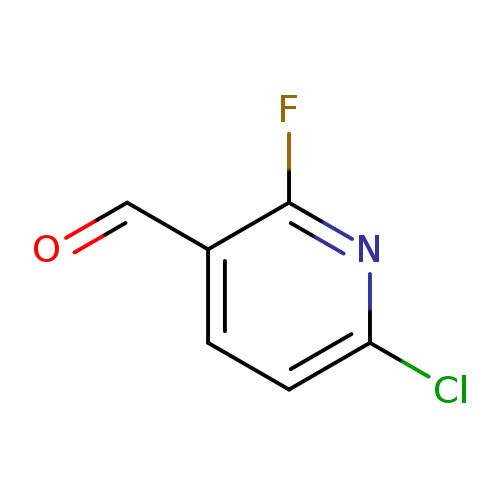
6-Chloro-2-fluoropyridine-3-carbaldehydeCatalog No.:AA003A8Y CAS No.:1093880-37-5 MDL No.:MFCD16556251 MF:C6H3ClFNO MW:159.5455 |
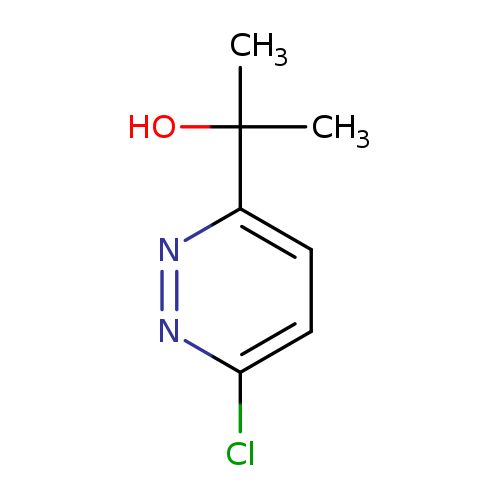
2-(6-Chloro-pyridazin-3-yl)-propan-2-olCatalog No.:AA0096DG CAS No.:1093881-08-3 MDL No.:MFCD16988471 MF:C7H9ClN2O MW:172.6122 |
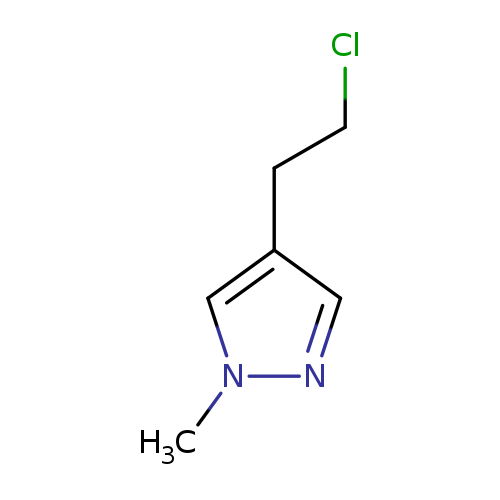
4-(2-Chloroethyl)-1-methyl-1h-pyrazoleCatalog No.:AA00HBDN CAS No.:1093881-63-0 MDL No.:MFCD16824702 MF:C6H9ClN2 MW:144.6021 |
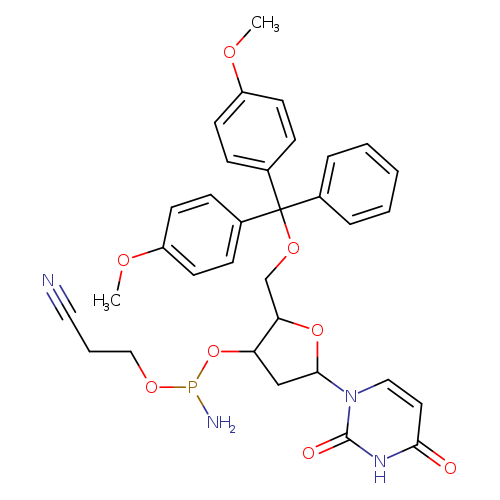
DMT-DU AMIDITE 0.25G, AB, SINGLECatalog No.:AA008UH1 CAS No.:109389-30-2 MDL No.:MFCD00036354 MF:C33H35N4O8P MW:646.6268 |
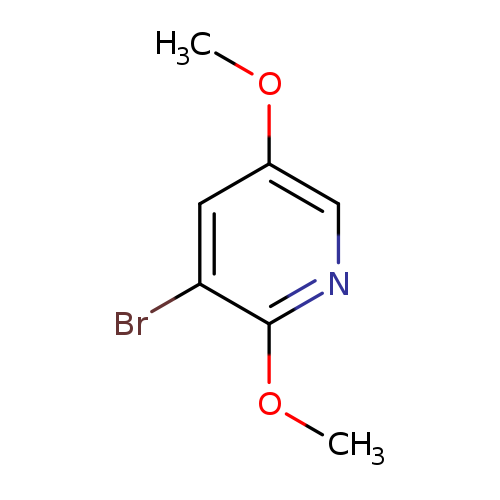
3-Bromo-2,5-dimethoxypyridineCatalog No.:AA00HBDP CAS No.:1093951-75-7 MDL No.:MFCD26398166 MF:C7H8BrNO2 MW:218.0479 |
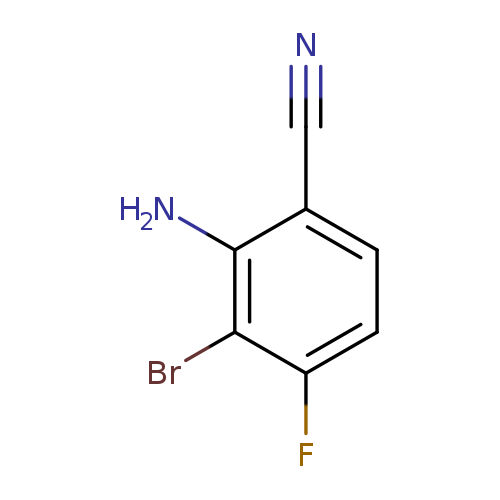
2-Amino-3-bromo-4-fluorobenzonitrileCatalog No.:AA0091XT CAS No.:1093951-76-8 MDL No.:MFCD18071063 MF:C7H4BrFN2 MW:215.0225 |
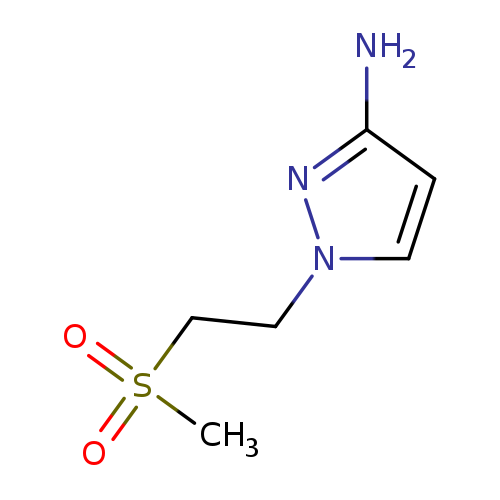
1-(2-methanesulfonylethyl)-1H-pyrazol-3-amineCatalog No.:AA01AKKI CAS No.:1093967-38-4 MDL No.:MFCD12806107 MF:C6H11N3O2S MW:189.2354 |
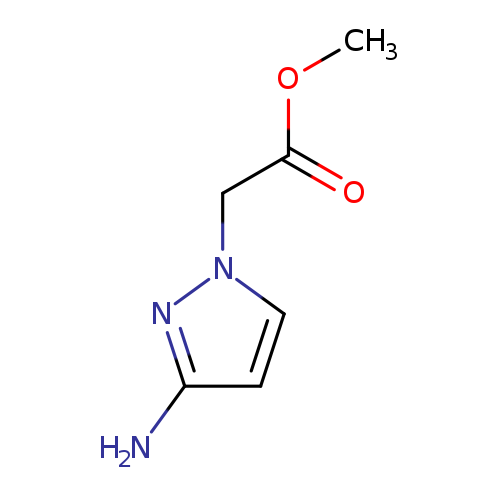
Methyl 2-(3-amino-1h-pyrazol-1-yl)acetateCatalog No.:AA00JDLK CAS No.:1093967-39-5 MDL No.:MFCD12149369 MF:C6H9N3O2 MW:155.1546 |
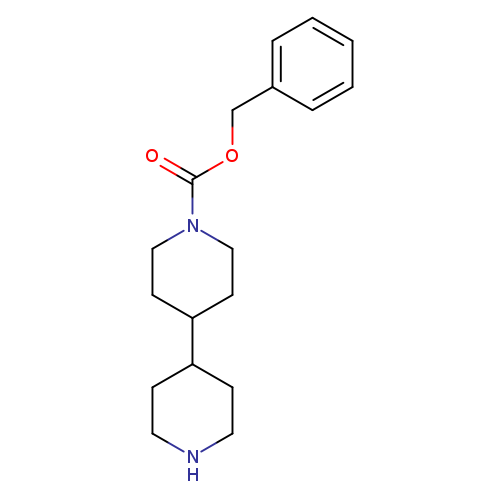
N-Cbz-4,4'-bipiperidineCatalog No.:AA007T3Q CAS No.:109397-72-0 MDL No.:MFCD02179170 MF:C18H26N2O2 MW:302.4112 |
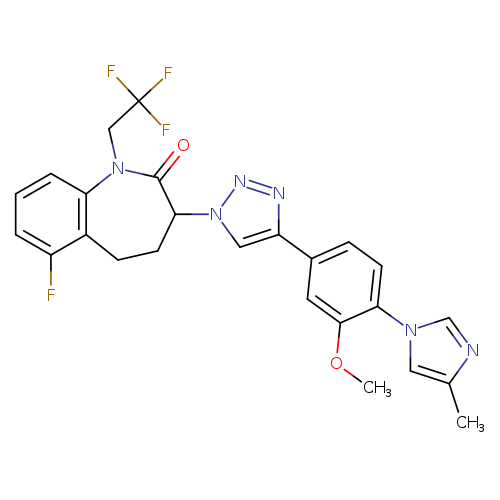
gamma-secretase modulator 2Catalog No.:AA008TDJ CAS No.:1093978-89-2 MDL No.:MFCD11977762 MF: MW: |
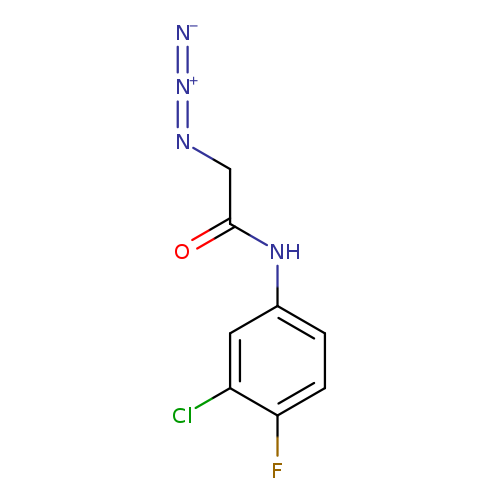
2-azido-N-(3-chloro-4-fluorophenyl)acetamideCatalog No.:AA00J0II CAS No.:1093981-49-7 MDL No.:MFCD14652443 MF:C8H6ClFN4O MW:228.6108 |
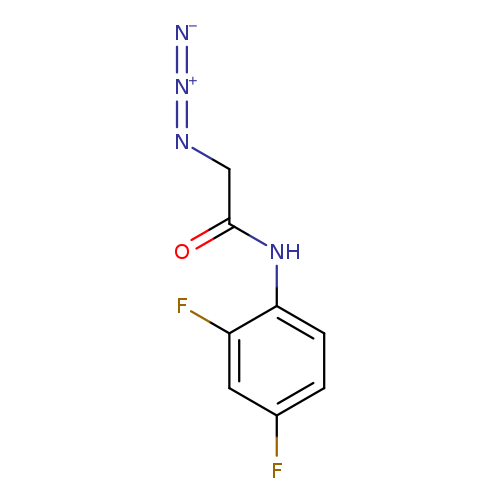
2-azido-N-(2,4-difluorophenyl)acetamideCatalog No.:AA00J0GD CAS No.:1093981-68-0 MDL No.:MFCD14652516 MF:C8H6F2N4O MW:212.1562 |
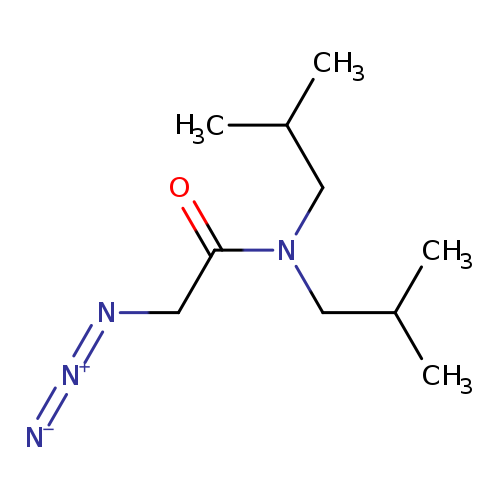
2-Azido-n,n-bis(2-methylpropyl)acetamideCatalog No.:AA01A15D CAS No.:1093981-79-3 MDL No.:MFCD14652432 MF:C10H20N4O MW:212.2920 |
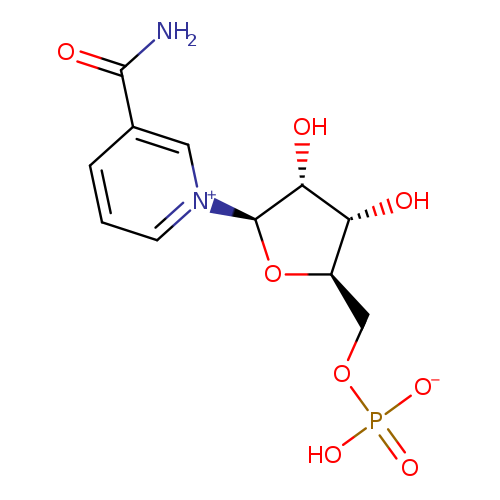
Beta-nicotinamide mononucleotideCatalog No.:AA003858 CAS No.:1094-61-7 MDL No.:MFCD00038748 MF:C11H15N2O8P MW:334.2192 |
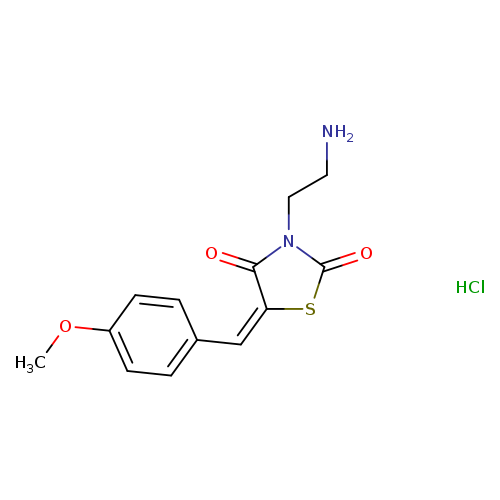
(5E)-3-(2-aminoethyl)-5-[(4-methoxyphenyl)methylidene]-1,3-thiazolidine-2,4-dione hydrochlorideCatalog No.:AA019GZO CAS No.:109401-52-7 MDL No.:MFCD08445261 MF:C13H15ClN2O3S MW:314.7878 |
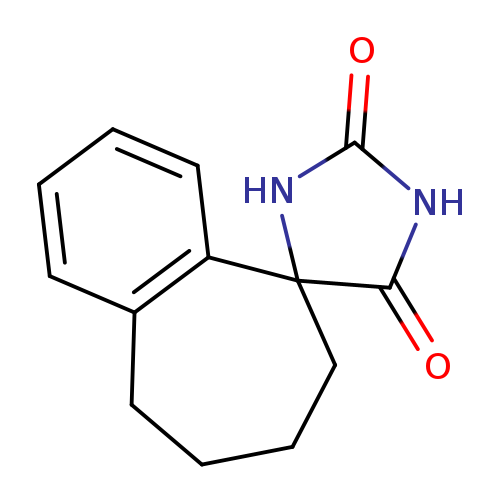
6,7,8,9-Tetrahydrospiro[benzo[7]annulene-5,4'-imidazolidine]-2',5'-dioneCatalog No.:AA0082DH CAS No.:109402-17-7 MDL No.:MFCD07838403 MF:C13H14N2O2 MW:230.2625 |
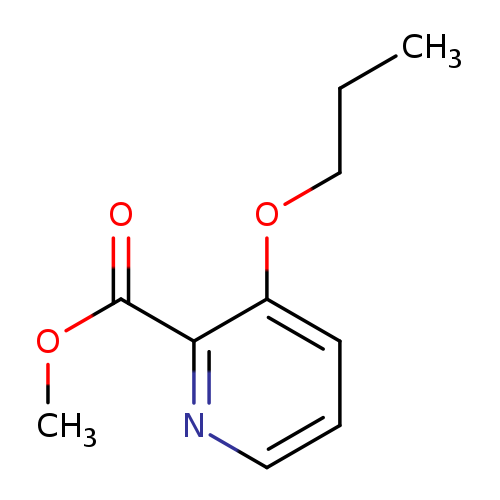
methyl 3-propoxypyridine-2-carboxylateCatalog No.:AA01BCYE CAS No.:1094025-00-9 MDL No.:MFCD28012377 MF:C10H13NO3 MW:195.2151 |
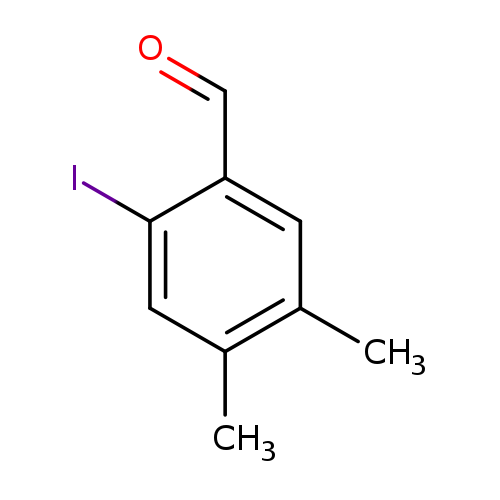
2-Iodo-4,5-dimethylbenzaldehydeCatalog No.:AA0096C9 CAS No.:1094026-85-3 MDL No.:MFCD14583067 MF:C9H9IO MW:260.0716 |
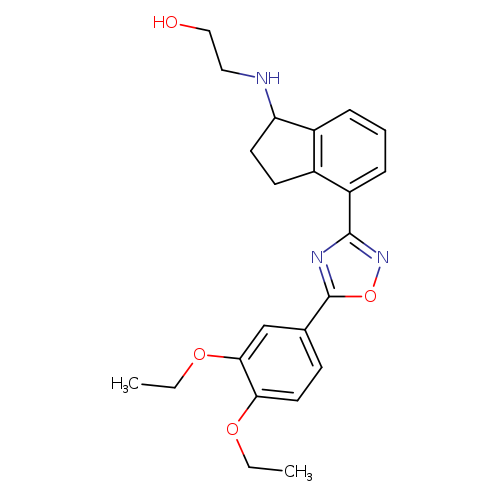
2-({4-[5-(3,4-diethoxyphenyl)-1,2,4-oxadiazol-3-yl]-2,3-dihydro-1H-inden-1-yl}amino)ethanol hydrochlorideCatalog No.:AA008SQI CAS No.:1094042-01-9 MDL No.:MFCD12912402 MF:C23H27N3O4 MW:409.4782 |
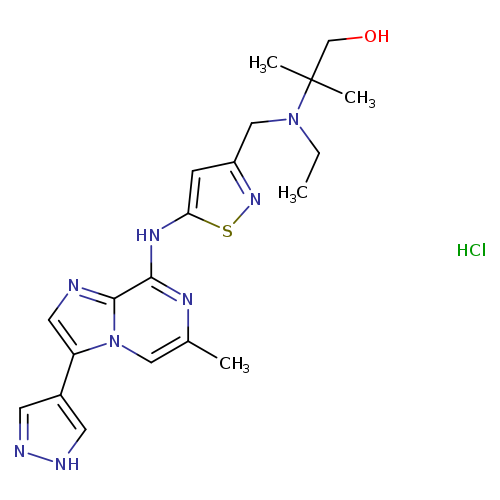
SCH-1473759 hydrochlorideCatalog No.:AA008TKC CAS No.:1094067-13-6 MDL No.:MFCD18251447 MF:C20H27ClN8OS MW:462.9994 |
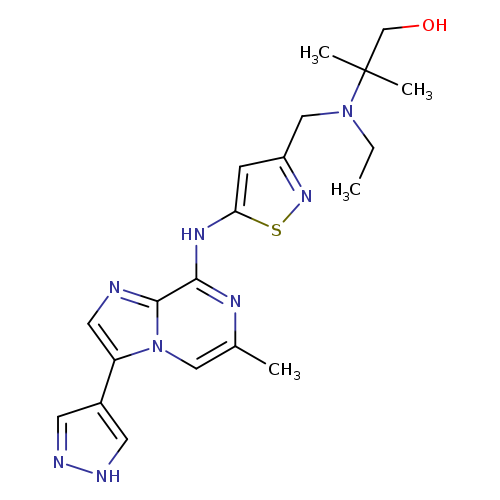
Sch-1473759Catalog No.:AA007AU7 CAS No.:1094069-99-4 MDL No.:MFCD18251446 MF:C20H26N8OS MW:426.5384 |
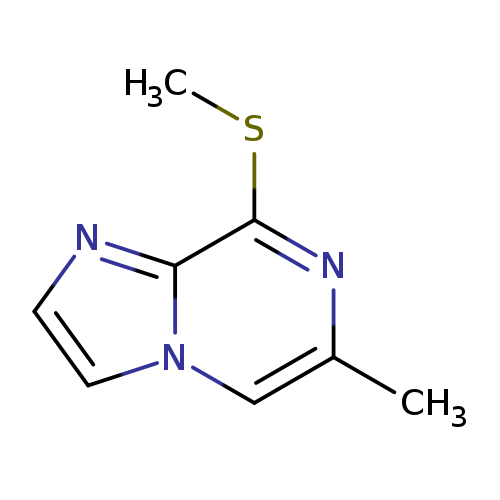
6-Methyl-8-methylsulfanyl-imidazo[1,2-a]pyrazineCatalog No.:AA0082DG CAS No.:1094070-46-8 MDL No.:MFCD17215754 MF:C8H9N3S MW:179.2422 |
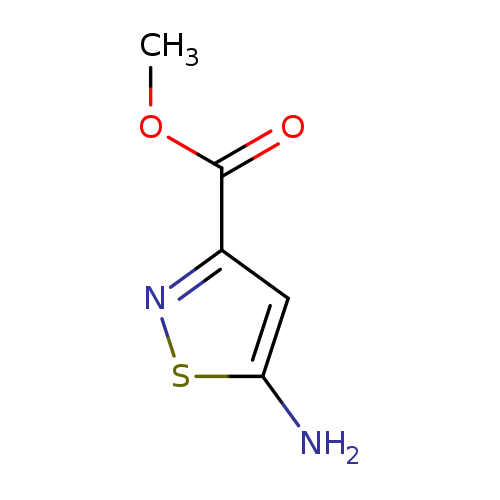
Methyl 5-aminoisothiazole-3-carboxylateCatalog No.:AA008YZV CAS No.:1094070-51-5 MDL No.:MFCD19105553 MF:C5H6N2O2S MW:158.1783 |
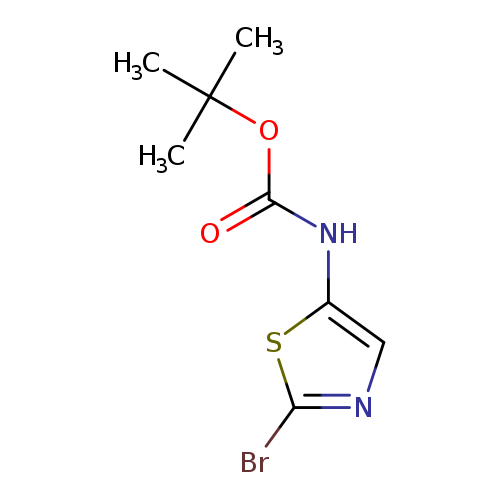
tert-Butyl 2-bromothiazol-5-ylcarbamateCatalog No.:AA0037VW CAS No.:1094070-77-5 MDL No.:MFCD13195442 MF:C8H11BrN2O2S MW:279.1541 |
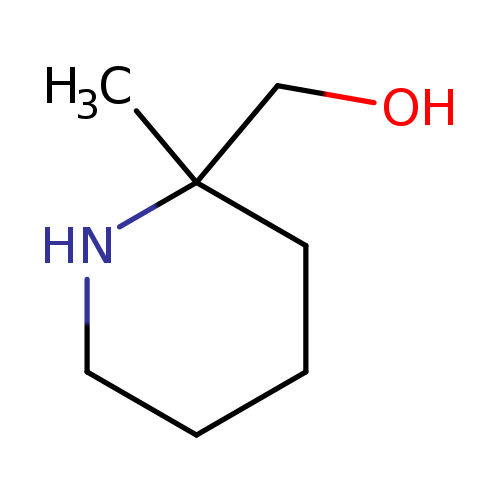
(2-Methylpiperidin-2-yl)methanolCatalog No.:AA00J35W CAS No.:1094071-26-7 MDL No.:MFCD14582623 MF:C7H15NO MW:129.2001 |
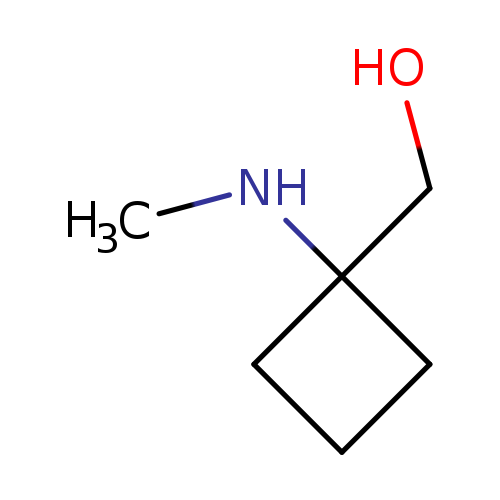
[1-(methylamino)cyclobutyl]methanolCatalog No.:AA01AL4E CAS No.:1094071-93-8 MDL No.:MFCD17170806 MF:C6H13NO MW:115.1735 |
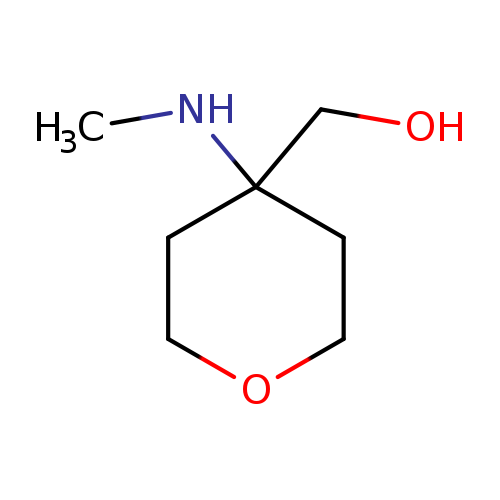
[4-(methylamino)oxan-4-yl]methanolCatalog No.:AA01BDCZ CAS No.:1094072-05-5 MDL No.:MFCD12826791 MF:C7H15NO2 MW:145.1995 |
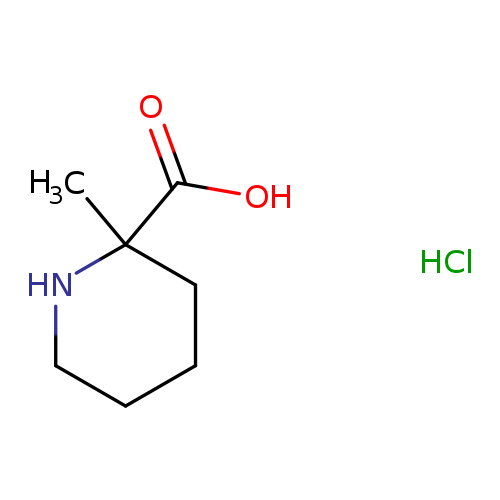
2-Methylpiperidine-2-carboxylic acid, HClCatalog No.:AA00IKSE CAS No.:1094072-12-4 MDL No.:MFCD19103470 MF:C7H14ClNO2 MW:179.6446 |
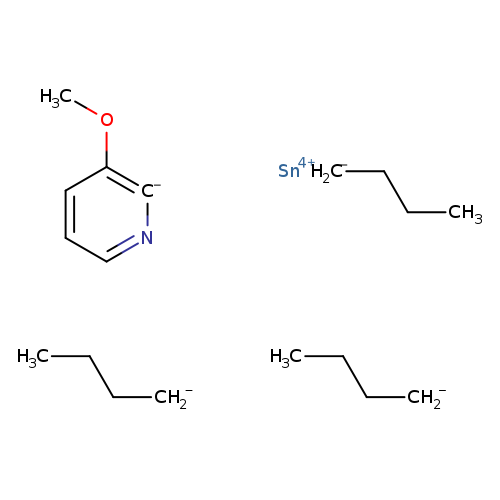
3-Methoxy-2-(tributylstannyl)pyridineCatalog No.:AA008TUO CAS No.:1094072-15-7 MDL No.:MFCD10699159 MF:C18H33NOSn MW:398.1617 |
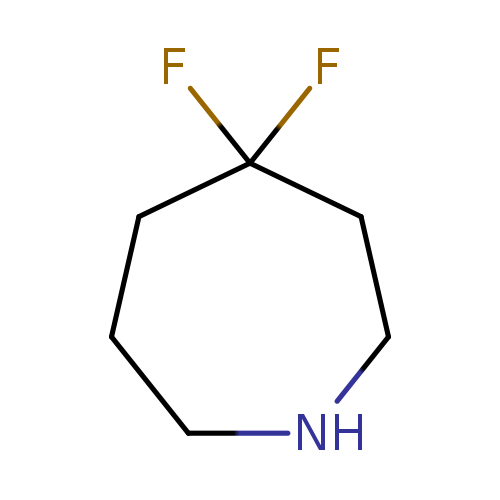
4,4-Difluoroazepane hydrochlorideCatalog No.:AA0097EZ CAS No.:1094073-72-9 MDL No.:MFCD11870210 MF:C6H11F2N MW:135.1550 |
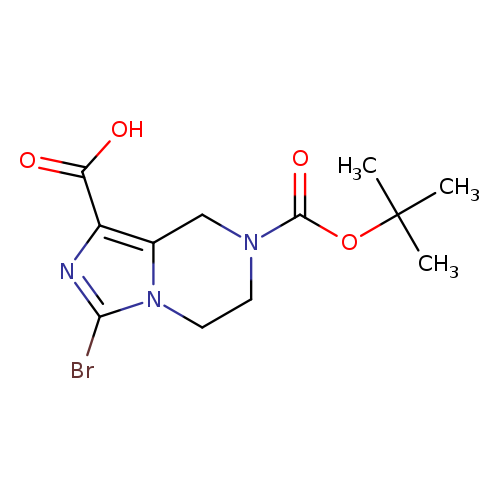
3-bromo-7-(tert-butoxycarbonyl)-5,6,7,8-tetrahydroimidazo[1,5-a]pyrazine-1-carboxylic acidCatalog No.:AA0091L6 CAS No.:1094091-46-9 MDL No.:MFCD22631518 MF:C12H16BrN3O4 MW:346.1771 |
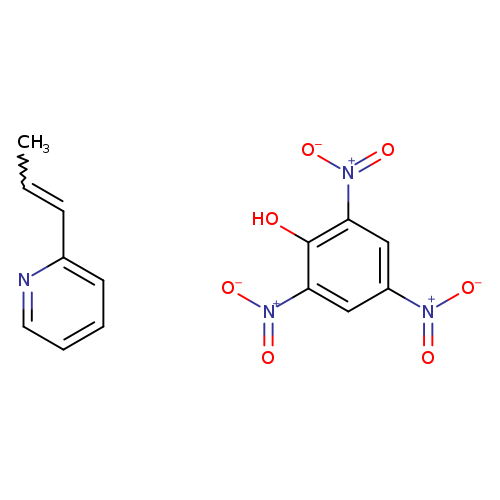
2,4,6-Trinitrophenol; 2-[(1E)-prop-1-en-1-yl]pyridineCatalog No.:AA01FG90 CAS No.:109410-54-0 MDL No.: MF:C14H12N4O7 MW:348.2677 |
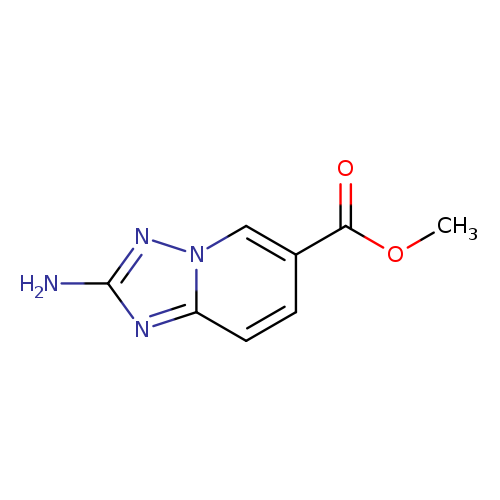
2-Amino-[1,2,4]triazolo[1,5-a]pyridine-6-carboxylic acid methyl esterCatalog No.:AA0093HA CAS No.:1094107-41-1 MDL No.:MFCD11976480 MF:C8H8N4O2 MW:192.1747 |
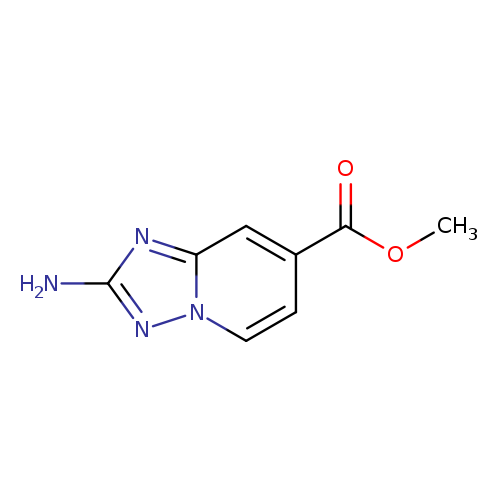
methyl 2-amino-[1,2,4]triazolo[1,5-a]pyridine-7-carboxylateCatalog No.:AA0093KS CAS No.:1094107-42-2 MDL No.:MFCD11975669 MF:C8H8N4O2 MW:192.1747 |
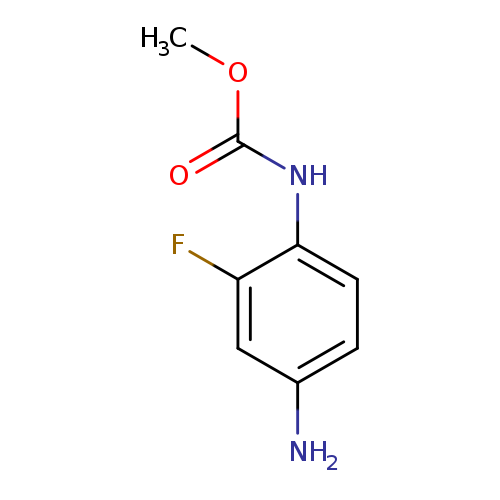
methyl N-(4-amino-2-fluorophenyl)carbamateCatalog No.:AA01A688 CAS No.:1094107-83-1 MDL No.:MFCD16074924 MF:C8H9FN2O2 MW:184.1677 |
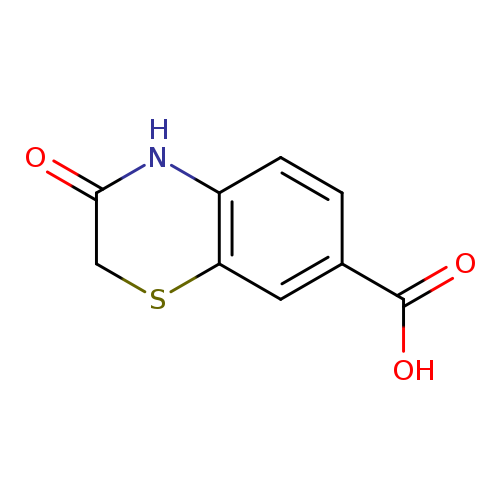
3-oxo-3,4-dihydro-2H-1,4-benzothiazine-7-carboxylic acidCatalog No.:AA01BBZ2 CAS No.:1094107-96-6 MDL No.:MFCD24484189 MF:C9H7NO3S MW:209.2218 |
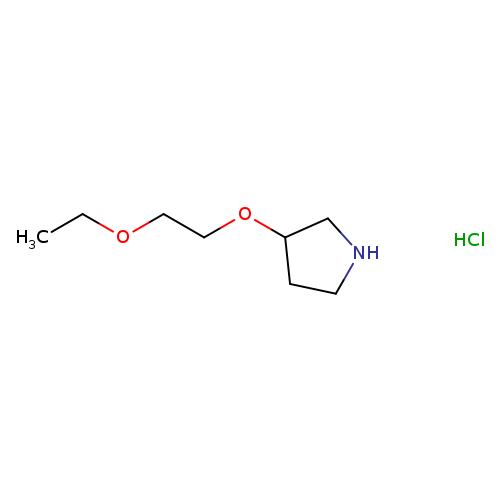
3-(2-ETHOXYETHOXY)PYRROLIDINE HYDROCHLORIDECatalog No.:AA009NRG CAS No.:1094108-02-7 MDL No.:MFCD09879227 MF:C8H18ClNO2 MW:195.6870 |
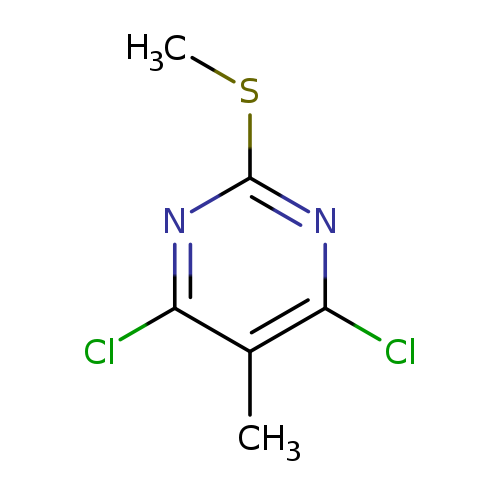
4,6-Dichloro-5-methyl-2-(methylthio)pyrimidineCatalog No.:AA00HBE0 CAS No.:109414-76-8 MDL No.:MFCD22417220 MF:C6H6Cl2N2S MW:209.0962 |
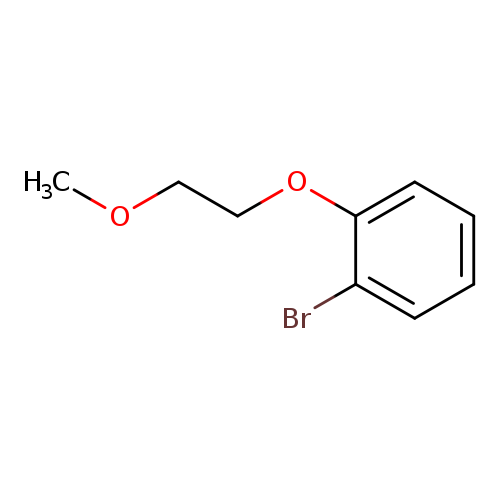
1-Bromo-2-(2-methoxyethoxy)benzeneCatalog No.:AA0082DC CAS No.:109417-60-9 MDL No.:MFCD09934824 MF:C9H11BrO2 MW:231.0864 |
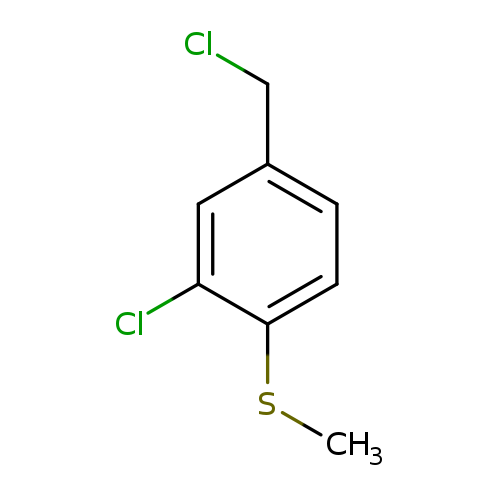
(2-Chloro-4-(chloromethyl)phenyl)(methyl)sulfaneCatalog No.:AA00926S CAS No.:109418-89-5 MDL No.:MFCD22489924 MF:C8H8Cl2S MW:207.1201 |
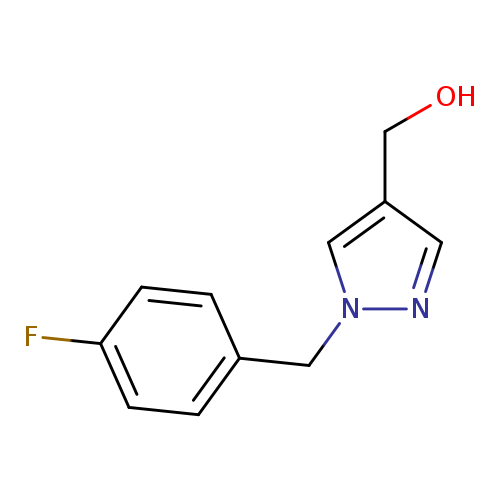
{1-[(4-fluorophenyl)methyl]-1H-pyrazol-4-yl}methanolCatalog No.:AA00HBE3 CAS No.:1094202-70-6 MDL No.:MFCD16862192 MF:C11H11FN2O MW:206.2162 |
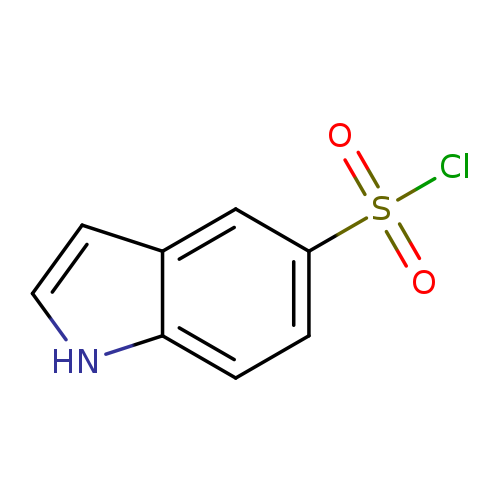
1H-Indole-5-sulfonyl chlorideCatalog No.:AA00911V CAS No.:1094209-33-2 MDL No.:MFCD19200539 MF:C8H6ClNO2S MW:215.6567 |
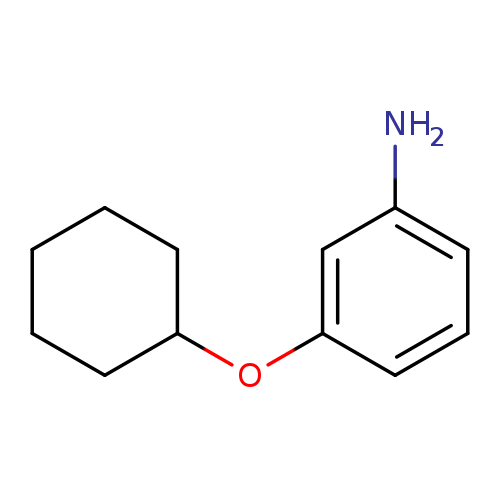
3-(cyclohexyloxy)anilineCatalog No.:AA01AADK CAS No.:109421-10-5 MDL No.:MFCD11167593 MF:C12H17NO MW:191.2695 |
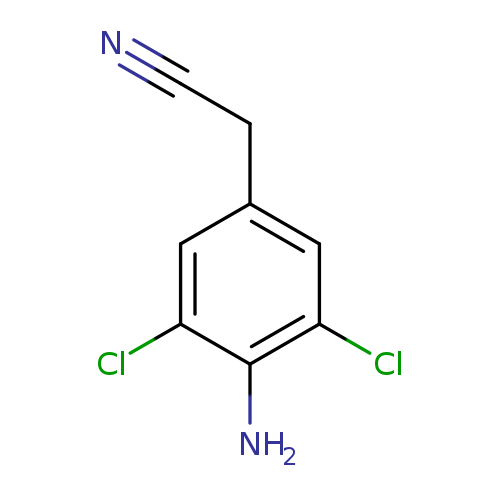
2-(4-Amino-3,5-dichlorophenyl)acetonitrileCatalog No.:AA01C3G6 CAS No.:109421-34-3 MDL No.:MFCD09991819 MF:C8H6Cl2N2 MW:201.0526 |
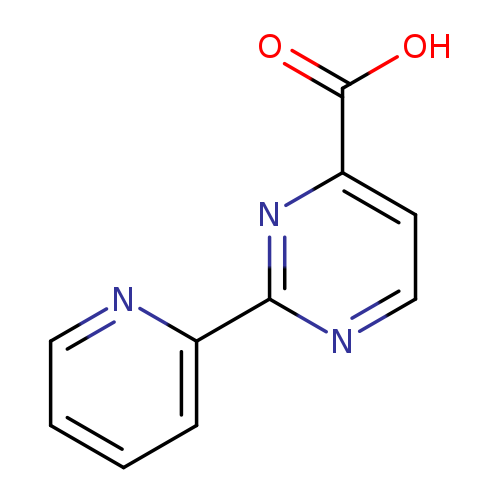
2-(Pyridin-2-yl)pyrimidine-4-carboxylic acidCatalog No.:AA0095QV CAS No.:1094211-77-4 MDL No.:MFCD16866579 MF:C10H7N3O2 MW:201.1815 |
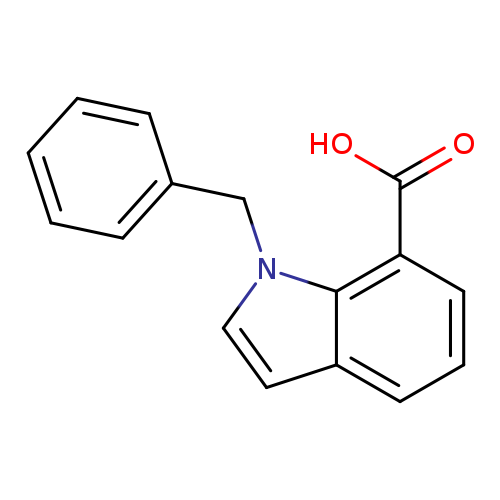
1-benzyl-1H-indole-7-carboxylic acidCatalog No.:AA01BJD0 CAS No.:1094217-58-9 MDL No.:MFCD11519414 MF:C16H13NO2 MW:251.2799 |
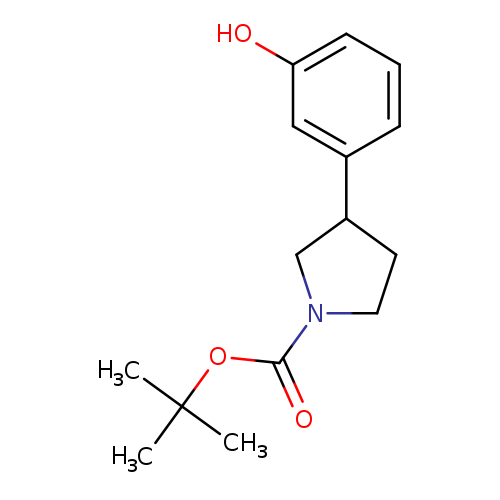
tert-Butyl 3-(3-hydroxyphenyl)pyrrolidine-1-carboxylateCatalog No.:AA008TL9 CAS No.:1094217-59-0 MDL No.:MFCD11519418 MF:C15H21NO3 MW:263.3321 |
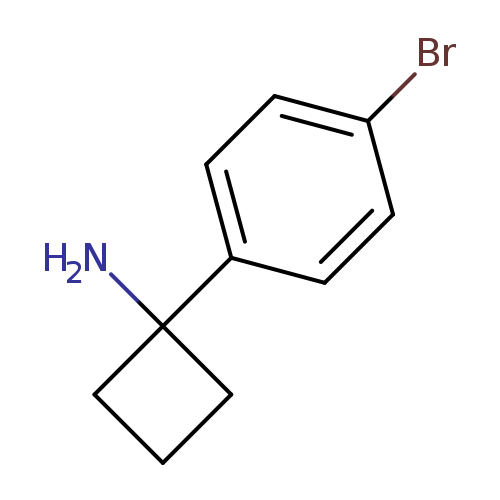
1-(4-Bromophenyl)cyclobutanamineCatalog No.:AA007T3M CAS No.:1094218-30-0 MDL No.:MFCD09910041 MF:C10H12BrN MW:226.1130 |
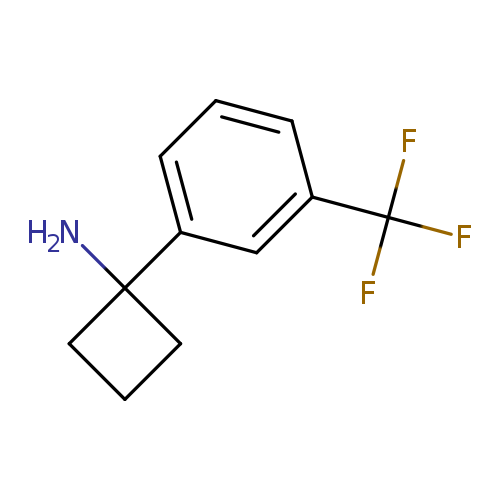
1-[3-(trifluoromethyl)phenyl]cyclobutan-1-amineCatalog No.:AA00HBE7 CAS No.:1094218-35-5 MDL No.:MFCD09864772 MF:C11H12F3N MW:215.2149 |
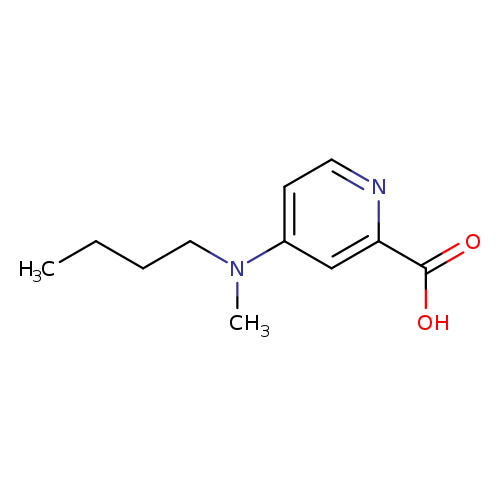
4-[Butyl(methyl)amino]pyridine-2-carboxylic acidCatalog No.:AA01AD3I CAS No.:1094219-12-1 MDL No.:MFCD11181459 MF:C11H16N2O2 MW:208.2569 |
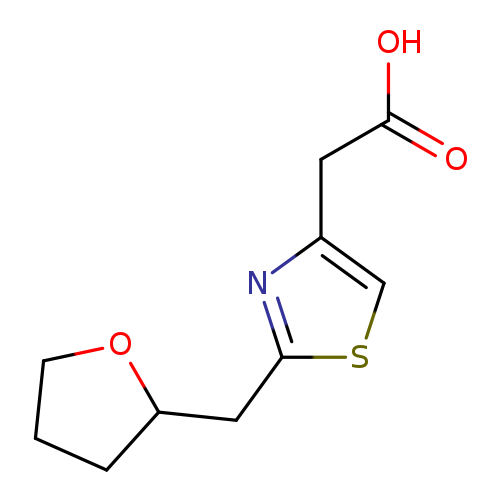
2-{2-[(oxolan-2-yl)methyl]-1,3-thiazol-4-yl}acetic acidCatalog No.:AA01C3D2 CAS No.:1094219-64-3 MDL No.:MFCD11180932 MF:C10H13NO3S MW:227.2801 |
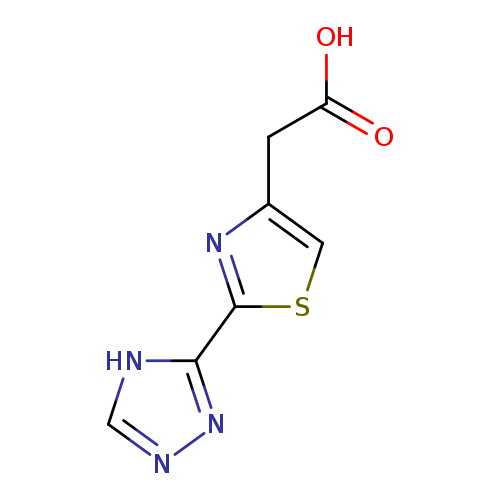
2-[2-(4H-1,2,4-triazol-3-yl)-1,3-thiazol-4-yl]acetic acidCatalog No.:AA01A9ZW CAS No.:1094219-65-4 MDL No.:MFCD11180936 MF:C7H6N4O2S MW:210.2131 |
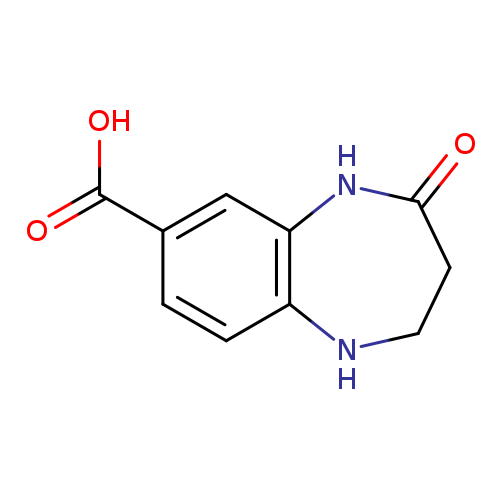
4-oxo-2,3,4,5-tetrahydro-1H-1,5-benzodiazepine-7-carboxylic acidCatalog No.:AA01AAEM CAS No.:1094219-66-5 MDL No.:MFCD11180938 MF:C10H10N2O3 MW:206.1980 |
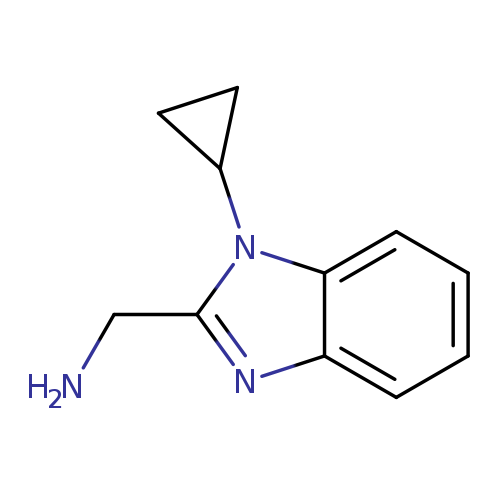
(1-cyclopropyl-1H-1,3-benzodiazol-2-yl)methanamineCatalog No.:AA01ABKQ CAS No.:1094220-55-9 MDL No.:MFCD11181233 MF:C11H13N3 MW:187.2410 |
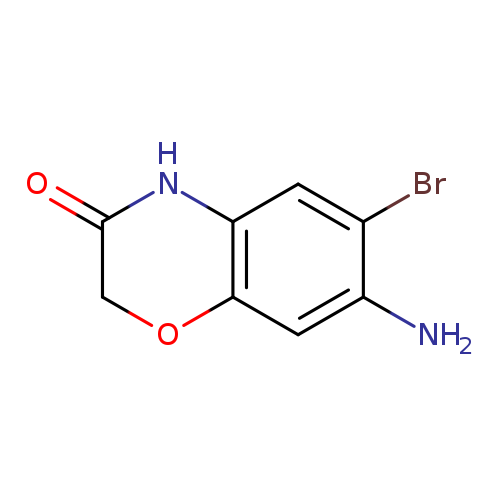
7-amino-6-bromo-3,4-dihydro-2H-1,4-benzoxazin-3-oneCatalog No.:AA019YP2 CAS No.:1094222-68-0 MDL No.:MFCD11205104 MF:C8H7BrN2O2 MW:243.0574 |
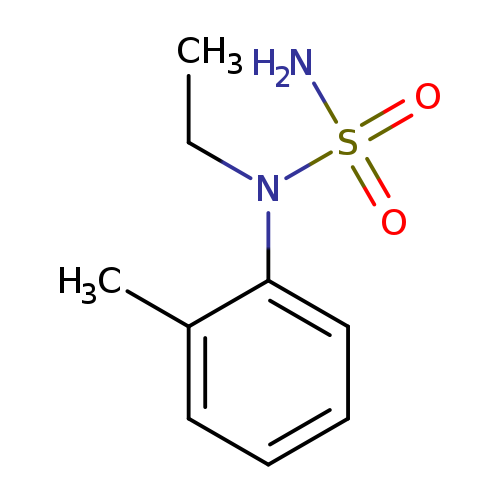
N-ethyl-N-(2-methylphenyl)aminosulfonamideCatalog No.:AA01B15E CAS No.:1094222-84-0 MDL No.:MFCD11205206 MF:C9H14N2O2S MW:214.2847 |
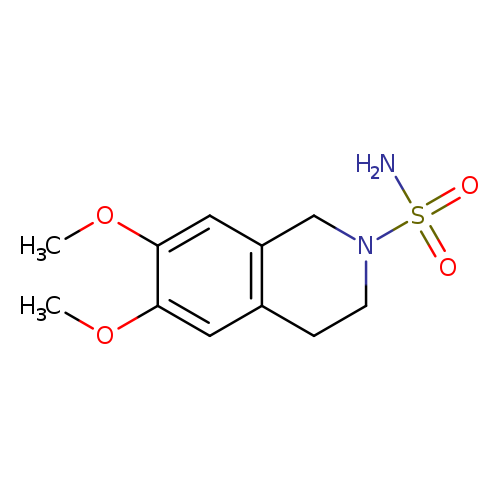
6,7-dimethoxy-1,2,3,4-tetrahydroisoquinoline-2-sulfonamideCatalog No.:AA01A8SM CAS No.:1094222-85-1 MDL No.:MFCD11205212 MF:C11H16N2O4S MW:272.3207 |
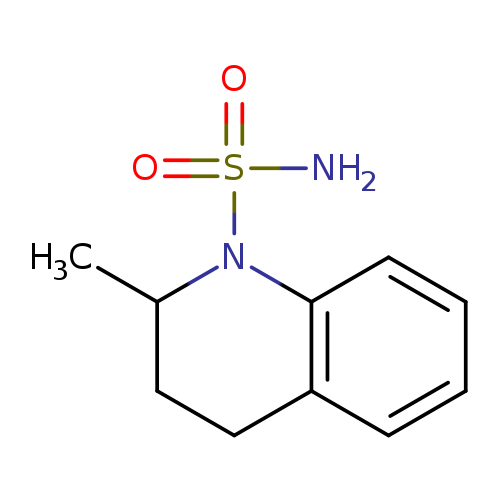
2-methyl-1,2,3,4-tetrahydroquinoline-1-sulfonamideCatalog No.:AA01A9R9 CAS No.:1094222-89-5 MDL No.:MFCD11205220 MF:C10H14N2O2S MW:226.2954 |
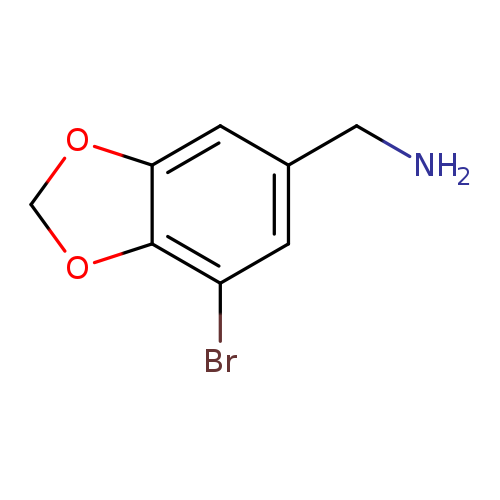
(7-bromo-2H-1,3-benzodioxol-5-yl)methanamineCatalog No.:AA019XL4 CAS No.:1094225-67-8 MDL No.:MFCD11182119 MF:C8H8BrNO2 MW:230.0586 |
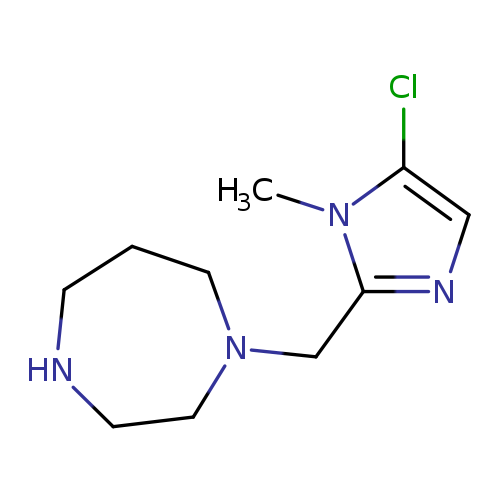
1-[(5-chloro-1-methyl-1H-imidazol-2-yl)methyl]-1,4-diazepaneCatalog No.:AA01AB8D CAS No.:1094226-29-5 MDL No.:MFCD11182309 MF:C10H17ClN4 MW:228.7218 |
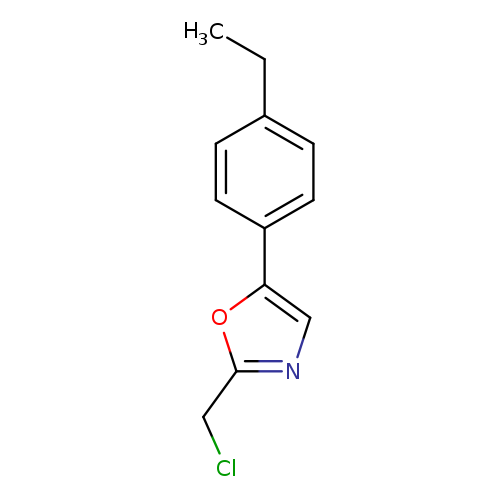
2-(chloromethyl)-5-(4-ethylphenyl)-1,3-oxazoleCatalog No.:AA01ELRP CAS No.:1094226-54-6 MDL No.:MFCD11182439 MF:C12H12ClNO MW:221.6828 |
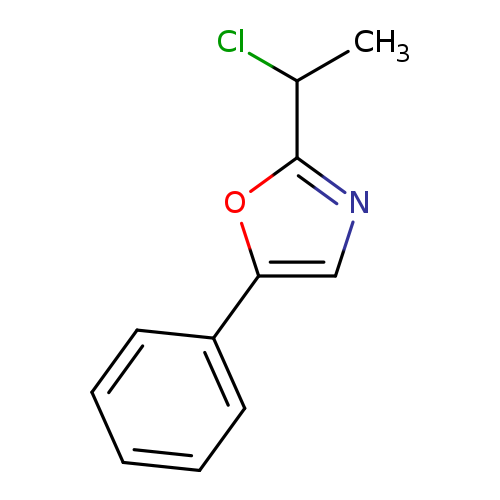
2-(1-Chloroethyl)-5-phenyl-1,3-oxazoleCatalog No.:AA01AIIQ CAS No.:1094226-56-8 MDL No.:MFCD11182446 MF:C11H10ClNO MW:207.6562 |
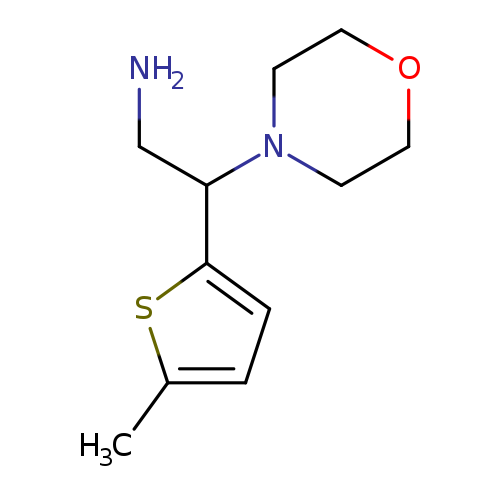
2-(5-methylthiophen-2-yl)-2-(morpholin-4-yl)ethan-1-amineCatalog No.:AA01AB8Y CAS No.:1094226-79-5 MDL No.:MFCD11186299 MF:C11H18N2OS MW:226.3384 |
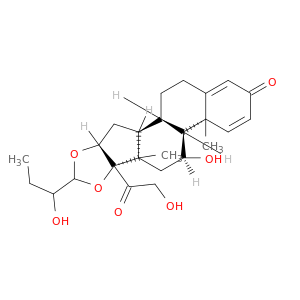
23-HydroxybudesonideCatalog No.:AA008W2Z CAS No.:109423-03-2 MDL No.: MF:C25H34O7 MW:446.5333 |
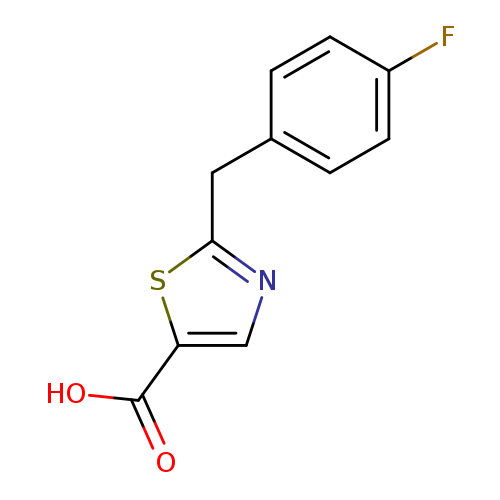
2-[(4-fluorophenyl)methyl]-1,3-thiazole-5-carboxylic acidCatalog No.:AA01A9Q9 CAS No.:1094230-04-2 MDL No.:MFCD11208428 MF:C11H8FNO2S MW:237.2501 |
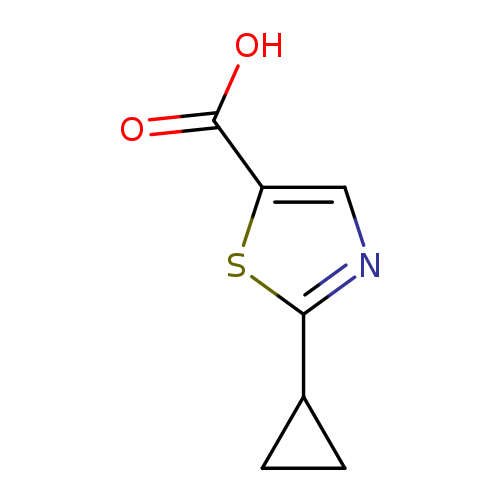
2-Cyclopropyl-1,3-thiazole-5-carboxylic acidCatalog No.:AA00998U CAS No.:1094230-05-3 MDL No.:MFCD11208434 MF:C7H7NO2S MW:169.2010 |
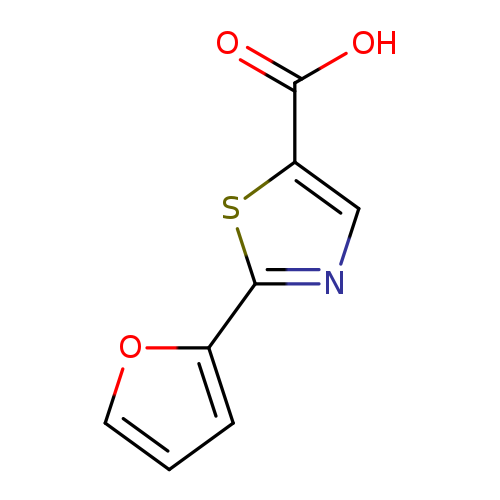
2-(furan-2-yl)-1,3-thiazole-5-carboxylic acidCatalog No.:AA01AAOD CAS No.:1094230-08-6 MDL No.:MFCD07376803 MF:C8H5NO3S MW:195.1952 |
3-cyclopentyl-3-oxo-2-phenylpropanenitrileCatalog No.:AA01AAW1 CAS No.:1094230-17-7 MDL No.:MFCD11208540 MF:C14H15NO MW:213.2750 |